| ID |
Date |
Author |
Subject |
|
181
|
Thu Aug 18 13:03:47 2022 |
Dylan Wells | Comparing inputs and outputs between AraSim and IceMC | Comparing Inputs:
Frequency Lists:
Ara - 83.33MHz - 1066.70 MHz, step = 13.33MHz
IceMC - 200MHz - 1500MHz, step = 10MHz
Number of files read in:
Ara - 1
IceMC - 8
Formating of files:
Ara - Theta, Phi, Gain (dB, thetra), Gain (theta), Phase (theta)
IceMc - Frequency, Gain (dB) (Different files cover different thetas and phis)
File Type:
Ara - .txt
IceMC - no suffix (file with 2 columns of text)
Other notes for IceMC inputs:
IceMC reads in 8 different files for gain.
vv_0 hh_0 vh_0 hv_0 vv_el vv_az hh_el hh_az
Found in ../anitaBuiltTool/components/icemc/data
vv_0 = gains for vertical polarization
hh_0 = gains for horizontal polarization
vh_0 = gains for v → h cross polarization
hv_0 = gains for h → v cross polarization
vv_el = v polarization, e angle
vv_az = v polarization, a angle
hh_el = h polarization, e angle
hh_az = h polarization a angle
for e angle and a angle in
0. 0
-
5
-
10
-
20
-
30
-
45
-
90
(iterates 1 to 6)
Comparing Outputs:
IceMC - veff is in second column in veff+runName+.txt file (in the outputDirectory directory)
Ara - veff is at the bottom of the AraOut.txt file |
|
Draft
|
Tue Jun 28 13:27:13 2022 |
Dylan Wells | Changes needed for the matching circuit script |
-
Fix the functions for the SLPC, SCPL, and PLSC L networks (change the paramaters to match with the format of our data)
-
Write the PCSL function
-
Create a function to find the number of L networks necessary (N) given a source and load resistance as well as a frequency range.
-
Write a function to broadband match two impedances given a source, load, central frequency, and N. (return a list of capacitances and inductances for the L networks)
|
|
243
|
Mon Jun 2 14:10:59 2025 |
Jacob Weiler | Building Status 06/02/2025 | We are almost to where we can start the physical building of the antenna!
I've attached all the information I currently have regarding the building project. Some of it is messy work notes and some is well-structured.
I’ve attached the following files for the GENETIS building project:
- Building Dump.txt
- My working notes that I used while trying to simulate the antenna in XFdtd (very messy)
- Building Dump of Useful Materials.txt
- List of materials that I found regarding the building project like slides, elogs, etc.
- Simulating Building Model.txt
- A writeup I made describing my process for simulating the antenna in XFdtd
- Done with change materials.zip
- Solidworks model of antenna
I also made a slide deck that contains the directory locations + has graphs HERE. |
| Attachment 1: Building_Dump_of_Useful_Materials.txt
|
Building Dump:
For Initial Building Run:
Generation 13, individual 84 seems to be result being used (this assumption is based on the fact that when trying to straighten the sides for building they used this individual)
/fs/ess/PAS1960/BiconeEvolutionOSC/BiconeEvolution/current_antenna_evo_build/XF_Loop/Evolutionary_Loop/Run_Outputs/2022_12_29
Elog Links for first building runs:
- Run Details: https://radiorm.physics.ohio-state.edu/elog/GENETIS/188
- Run Results + Gain Patterns: https://radiorm.physics.ohio-state.edu/elog/GENETIS/189
- Matching Circuit PCB: https://radiorm.physics.ohio-state.edu/elog/GENETIS/193
- Matching Circuit Parts: https://radiorm.physics.ohio-state.edu/elog/GENETIS/191
- Matching Circuit Schematic: https://radiorm.physics.ohio-state.edu/elog/GENETIS/230
- Matching Circuit Initial Design: https://radiorm.physics.ohio-state.edu/elog/GENETIS/183
- PoR Plots 1: https://radiorm.physics.ohio-state.edu/elog/GENETIS/194
- PoR Plots 2: https://radiorm.physics.ohio-state.edu/elog/GENETIS/196
- Straightened Sides 1: https://radiorm.physics.ohio-state.edu/elog/GENETIS/229
- Straightened Sides 2: https://radiorm.physics.ohio-state.edu/elog/GENETIS/236
- Engineering Call: https://docs.google.com/presentation/d/1Lo_6mFTmPbkToTrEeOpvznSdbxyPZexPag1qijTeYyM/edit?usp=sharing
At some point for some reason, another run seems to have been created for building with the crazy sides run here:
/fs/ess/PAS1960/BiconeEvolutionOSC/BiconeEvolution/current_antenna_evo_build/XF_Loop/Evolutionary_Loop/Run_Outputs/2023_09_05_realized_curved_run
Top 5 vEffective Scores:
Value: 5.09897, Generation: 41, Individual: 44 (Seems to be this one, modified)
Value: 5.07746, Generation: 37, Individual: 16
Value: 5.05508, Generation: 37, Individual: 5
Value: 5.04558, Generation: 38, Individual: 12
Value: 5.04026, Generation: 48, Individual: 5
GENETIS Useful Links:
- GENETIS Google Drive: https://drive.google.com/drive/folders/1iDamk46R2_oOLHtvsOg4jNy05mCiB7Sn?dmr=1&ec=wgc-drive-hero-goto
- Onboarding Materials: https://radiorm.physics.ohio-state.edu/elog/GENETIS/41
- Julie's Dissertation: https://radiorm.physics.ohio-state.edu/elog/Write-Ups/220404_161525/Julie_Rolla_Dissertation.pdf
- Julie's Candidacy: https://as-phy-radiorm.asc.ohio-state.edu/elog/Write-Ups/44
- ICRC Proceedings: https://arxiv.org/pdf/2112.00197
- Phys Rev D Paper: https://journals.aps.org/prd/abstract/10.1103/PhysRevD.108.102002
- ARA Loop GitHub: https://github.com/osu-particle-astrophysics/GENETIS-ARA
- PUEO Loop GitHub: https://github.com/osu-particle-astrophysics/GENETIS_PUEO
- Shared Code GitHub: https://github.com/osu-particle-astrophysics/Shared-Code
- AraSim GitHub: https://github.com/ara-software/AraSim/tree/master
- pueoSim GitHub: https://github.com/PUEOCollaboration/pueoSim
|
| Attachment 2: Simulating_Building_Model.txt
|
Simulating Building Model
Getting the model we want to build from Solidworks into XF ready for simulations took a bit of work. Here are the steps and things I did to get it to finally work with materials and everything enabled (minus conductors in the coax cable).
General Instructions to setup the antenna the same as I did. Saving after each of these steps.
Getting out of Solidworks:
To get out of Solidworks, I used .step file under the assumption that it would carry the material data over into XF (this assumption was based on what I had read online, though I was looking at the wrong places for that information as I found out later). With this assumption, we spent time getting the materials correct in Solidworks before exporting out into the .step file. I spent considerable time double checking the materials in Solidworks to make sure that everything was defined correctly with at least good enough approximations of the materials to get a simulation working.
Material definitions:
Shells: Plastic wrapped in Copper Foil, approximated by just having the whole shell as Copper
Screws Connecting horizontal halves: ABS (PEEK Plastic)
Supports Connecting vertical halves: ABS
Other screws: Non-magnetic stainless steel (Passivated 18-8 Stainless Steel)
Coax Cable:
Dielectric: Foam Polyethylene (FPE)
Inner Conductor: Solid Bare Copper Covered Aluminum
Outer Conductor: Aluminum Tape
Outer Braid: Tinned Copper
Jacket: Polyethylene
Everything else: Approximated as copper (not entirely sure if they are copper fully or if they are just wrapped with copper foil)
Importing into XFdtd:
After having the step file exported, I put the file onto OSC and opened a new XFdtd project. I then clicked the "Import -> Cad Models" to select my file and have it imported. I did not import in the material data as I found out it did not import in correctly to each part, so I ignored it and manually added the material definitions later.
I now have the model into XFdtd, but it’s rotated 90 degrees to be in the horizontal plane. This isn’t inherently bad, but I want my surrounding scripts to not have to be changed much so I rotate the model to have the wire pointing in the +z direction in XFdtd. Once I’ve done this, I right click on the Braid and Inner + Outer conductors in the coax cable and select something similar to "Do not include in meshing." This now makes sure that these are NOT in the simulation.
Then, I manually added material properties into XFdtd from definitions I looked up online for the electric + magnetic properties of:
Copper
Plastic (ABS)
Foam Polyethylene (dielectric)
Polyethylene
Aluminum
Stainless Steel
After creating these material definitions, I applied them to the appropriate parts.
Feed adjustments:
I want the feed to be in the same location as the coax cable for the best results, problem is that there are holes in the place where the coax cable would be split (which I disabled to prevent shorting!). So, I setup two copper pucks (not much thicker than the copper pieces that cover the tops by the feeds) to fill out the holes and make sure each half is connected to each corresponding side of the feed. After I place these in the correct location, I use the same 50-ohm feed setup script used in the GENETIS Vpol loop.
Now we have everything almost ready to simulate.
Simulation Setup:
There are various things needed to be done to setup the script, and while you can use the GUI, I’m not familiar enough with it so I just used the corresponding scripts in the GENETIS loop that would be needed before an XF simulation takes place. After I run this, we are now ready to simulate.
Running Simulation:
Again, not familiar with the GUI so I just used the GENETIS XFdtd job scripts and modified them for this purpose (which was just adjusting directories of outputs). Then I submitted the job and waited for it to complete (I believe it took around 8 min per simulation for this antenna).
Getting uan files:
I then opened the simulation and ran the same code used to output UANs as used in the GENETIS loop to output all 60 uan files at the frequencies we want.
Now you should have the files for the building model that was made in CAD!
Debugging Steps I took:
This took me a while over spring break, at least a lot longer than I thought it would.
I found out that the material data from .step file does not translate as I had expected into XFdtd so I had to manually input the material data as shown above
I found out that hiding a part in XF does NOT exclude it from simulation, you have to remove it from meshing or it still remains there
I did compare the geometries between the as-evolved antenna and this building model, there are differences but they are slight. Overall they are very similar
Removing the conductors for the coax cable is necessary as it will just short the two pieces (leading back to 2) which makes sense
Final:
After doing all this, I ended up getting what I deemed reasonable for the outputs for the building model after 28 runs in my 03_13_2025_manual.xf xf file on my user. Run28 is the run that I describe setting up above this text.
The material is not 1-1 with what will be built as I found it difficult to find exact electro-magnetic properties for all of these, so maybe the discrepancies in gain could be resolved through more rigorous definitions. It could actually technically make it worse, but maybe when this is physically built this will need to be done to get more accurate results to compare against.
Simulating both of these with higher statistics in AraSim resulted in the antennas actually performing worse than the base Vpol antenna, which stinks but it is both of the antennas not just one!
After (delayed) emails back and forth with Christian Miki from University of Hawaii, he found these same issues while he was looking at the model from CAD before I went through the XFdtd simulation steps.
Material Definitions in XF:
For critique, here are the material definitions I used in the XF simulation (using XF material definition windows). You should be able to look at them in the actual xf project I mentioned above in my user (full path in slide decks)
All setup with the following:
Type: Physical
Electric: Isotropic
Magnetic: Isotropic
Passivated 18-8 Stainless Steel:
Electric Tab
Type: Nondispersive
Entry Method: Normal
Good Conductor: Automatic
Conductivity: 1.1e+06 S/m
Relative Permittivity: 1
Infinite Dielectric Strength: Yes
Magnetic Tab
Type: Nondispersive
Entry Method: Normal
Conductivity: 0
Relative Permeability: 1.03
PEEK Plastic
Electric Tab
Type: Nondispersive
Entry Method: Loss Tangent
Good Conductor: Automatic
Relative Permittivity: 3.3
Loss Tangent: 0.003
Evaluation Frequency: 1 MHz
Infinite Dielectric Strength: Yes
Magnetic Tab
Type: Nondispersive
Entry Method: Normal
Conductivity: 0
Relative Permeability: 1
Foam Polyethylene
Electric Tab
Type: Nondispersive
Entry Method: Loss Tangent
Good Conductor: Automatic
Relative Permittivity: 1.6
Loss Tangent: 0.0004
Evaluation Frequency: 1 MHz
Infinite Dielectric Strength: Yes
Magnetic Tab
Type: Nondispersive
Entry Method: Normal
Conductivity: 0
Relative Permeability: 1
Polyethylene
Electric Tab
Type: Nondispersive
Entry Method: Loss Tangent
Good Conductor: Automatic
Relative Permittivity: 2.25
Loss Tangent: 0.0004
Evaluation Frequency: 1 MHz
Infinite Dielectric Strength: Yes
Magnetic Tab
Type: Nondispersive
Entry Method: Normal
Conductivity: 0
Relative Permeability: 1
ABS Plastic
Electric Tab
Type: Nondispersive
Entry Method: Loss Tangent
Good Conductor: Automatic
Relative Permittivity: 3.2
Loss Tangent: 0.005
Evaluation Frequency: 1 MHz
Infinite Dielectric Strength: Yes
Magnetic Tab
Type: Nondispersive
Entry Method: Normal
Conductivity: 0
Relative Permeability: 1
Copper Foil
Electric Tab
Type: Nondispersive
Entry Method: Normal
Good Conductor: Automatic
Conductivity: 5.96e+07 S/m
Relative Permittivity: 1
Infinite Dielectric Strength: Yes
Magnetic Tab
Type: Nondispersive
Entry Method: Normal
Conductivity: 0
Relative Permeability: 1
|
| Attachment 3: Building_Dump.txt
|
Building Dump:
Debugging Issues with Antenna model simulation:
Graphs to get (compared to Curved_Sides Antenna Run):
- Gain Plots
- Look at frequencies where dips. Could be due to: dielectric loss, mismatched impedance or structural changes
- Impedance Over Frequency Plots
- Want impedance to be around 50 Ohm for resistive components and 0 for reactance at operational frequencies
- S11 Plots (Return Loss VS Frequency)
- Look for where the S11 dips to determine where the antenna is resonant
- Total Efficiency Vs Frequencies
- Drops at certain frequencies indicates problems!
- VSWR vs Frequency
- Lower VSWR means better matching
in 03_13_2025_manual.xf:
Run1 = wrong material defs (deleted)
Run2 = glitched it
Run3 = wrong material def again slightly changed tho
Run4 = wrong material, with conductor gone
Run5 = wrong material, with full wire gone
Run6 = right material, full wire gone
Run7 = right material, conductor gone
Run8 = right material, everything there feed shifted to side
Run9 = right material, feed in middle of conductor
Run10 = right material, wire gone feed offset reduced (putting closer to center). this failed because the top of the feed was disconnected
Run11 = right material, feed with correct max feed offset allowed, coax gone
Run12 = trying the same thing but with the coax gone with building the feed
Run13 = with coax cable back, feed shifted closer to middle (apparently forgot to save and it's just the same thing.. as run12)
Run14 = adding pads and putting feed in the middle of the antenna, leave dielectric and jacket turned on
Run16 = pads, feed in middle, removing dielectric and jacket (-300 thing again.. not sure why)
Run17 = same thing but ABS material changed and adjusted pucks a little
Run18 = same ABS material change but with only inner conductor removed (I am testing why I am getting -300..)
Run19 = removed supports, still with pucks + only inner conductor removed
Run20 = removing pucks, with conductor removed and new ABS material (no more -300 but very low again...)
run21 = removed pucks, conductors(PLURAL) with new ABS Material
run23 = back to just supports, offset feed new ABS Material
run24 = coax gone, og ABS material, with the offset feed closer to the middle
run25 = everything back to normal coax gone (something wrong)
run26 = trying to fix the issue I'm seeing (FIXED) you have to uncheck that materials are included in meshing :/
run27 = actually removing the outer and inner conductors (yields worse gains!)
run28 = moving to feed center w/ copper plates and with the jacket + dielectric
Seems like the wire in the middle should be plastic (or non-conducting)? based off document wangjie sent me
"If we 3D print the metal, Chi-Chih thought that we could keep them together through a plasic rod running
through the middle" (It's not!)
maybe not, named LMR600 in solidworks which have the following material properties:
https://www.awcwire.com/lmr-cable/lmr-75-ohm-cable/lmr-600-75
screws connecting halves needed to be plastic
all other screws needed to be non-magnetic stainless steel
everything else is copper (?)
Trying to change materials of the wire and supports (03_13_2025_building_sim_2.xf): still bad
Trying again with same materials and putting feed down center of coax cable(03_13_2025_building_sim_3.xf): everything is -300 dBi :(
removing the copper middle part (03_13_2025_building_sim_4.xf): still bad
manually adding materials into XFdtd (03_13_2025_manual.xf): still bad, but different bad actually numbers-wise worse
- Passivated 18-8 Stainless Steel
- PEEK Plastic
- Dielectric: Foam Polyethylene (FPE)
- Inner Conductor: Solid Bare Copper Covered Aluminum
- Outer Conductor: Aluminum Tape
- Outer Braid: Tinned Copper
- Jacket: Polyethylene
- ABS Plastic
- Copper foil
I believe the feed replaces the coax cable in the middle so I am removing the inner conductor and assuming that it will be the same as the feed.
Dimensions of Curved Antenna (model based off this): (in cm for relevant parts)
- r1 = 3.20675
- height1 = 39.3683
- a1 = -0.0123505
- b1 = 0.418171
- r2 = 3.6116
- height2 = 18.605
- a2 = -0.0233028
- b2 = 0.369081
- Total height = 60.9733
Dimensions of Model in XF: (ignoring a's and b's as that's harder to measure..) (again in cm) (rough measurements in XF)
- r1 = 3.7
- h1 = 33.7441
- r2 = 3.4
- h2 = 18.71
- total height = 55.45 (no cable) 60.6459 (including cable)
Reference run XF settings:
- Removed the wire in the middle that was connecting the two sides: no difference (need to redo with it actually deleted + having the top plates copper) (03_11_2025_building_sim_1.xf)
- Removed middle wire AGAIN (03_13_2025_building_sim_0.xf): no difference, same issue
- Removed Supports and simulated(03_12_2025_building_sim_0.xf): This seems to have fixed the issue I'm seeing, so either the supports or the wire are shorting the antenna (or both!)
- Removed Supports ONLY(03_13_2025_building_sim_1.xf): still happening, though less extreme
For Initial Building Run:
Generation 13, individual 84 seems to be result being used (this assumption is based on the fact that when trying to straighten the sides for building they used this individual)
/fs/ess/PAS1960/BiconeEvolutionOSC/BiconeEvolution/current_antenna_evo_build/XF_Loop/Evolutionary_Loop/Run_Outputs/2022_12_29
Elog Links for first building runs:
- Run Details: https://radiorm.physics.ohio-state.edu/elog/GENETIS/188
- Run Results + Gain Patterns: https://radiorm.physics.ohio-state.edu/elog/GENETIS/189
- Matching Circuit PCB: https://radiorm.physics.ohio-state.edu/elog/GENETIS/193
- Matching Circuit Parts: https://radiorm.physics.ohio-state.edu/elog/GENETIS/191
- Matching Circuit Schematic: https://radiorm.physics.ohio-state.edu/elog/GENETIS/230
- Matching Circuit Initial Design: https://radiorm.physics.ohio-state.edu/elog/GENETIS/183
- PoR Plots 1: https://radiorm.physics.ohio-state.edu/elog/GENETIS/194
- PoR Plots 2: https://radiorm.physics.ohio-state.edu/elog/GENETIS/196
- Straightened Sides 1: https://radiorm.physics.ohio-state.edu/elog/GENETIS/229
- Straightened Sides 2: https://radiorm.physics.ohio-state.edu/elog/GENETIS/236
At some point, another run seems to have been created for building with the crazy sides run here with REALIZED GAIN:
/fs/ess/PAS1960/BiconeEvolutionOSC/BiconeEvolution/current_antenna_evo_build/XF_Loop/Evolutionary_Loop/Run_Outputs/2023_09_05_realized_curved_run
- Run is using the same freq of interest as what we currently use!!
Top 5 vEffective Scores of Realized Gain run:
Value: 5.09897, Generation: 41, Individual: 44 (Seems to be this one, modified)
Value: 5.07746, Generation: 37, Individual: 16
Value: 5.05508, Generation: 37, Individual: 5
Value: 5.04558, Generation: 38, Individual: 12
Value: 5.04026, Generation: 48, Individual: 5
GENETIS Useful Links:
- GENETIS Google Drive: https://drive.google.com/drive/folders/1iDamk46R2_oOLHtvsOg4jNy05mCiB7Sn?dmr=1&ec=wgc-drive-hero-goto
- Onboarding Materials: https://radiorm.physics.ohio-state.edu/elog/GENETIS/41
- Julie's Dissertation: https://radiorm.physics.ohio-state.edu/elog/Write-Ups/220404_161525/Julie_Rolla_Dissertation.pdf
- Julie's Candidacy: https://as-phy-radiorm.asc.ohio-state.edu/elog/Write-Ups/44
- ICRC Proceedings: https://arxiv.org/pdf/2112.00197
- Phys Rev D Paper: https://journals.aps.org/prd/abstract/10.1103/PhysRevD.108.102002
- ARA Loop GitHub: https://github.com/osu-particle-astrophysics/GENETIS-ARA
- PUEO Loop GitHub: https://github.com/osu-particle-astrophysics/GENETIS_PUEO
- Shared Code GitHub: https://github.com/osu-particle-astrophysics/Shared-Code
- AraSim GitHub: https://github.com/ara-software/AraSim/tree/master
- pueoSim GitHub: https://github.com/PUEOCollaboration/pueoSim
|
| Attachment 4: done_with_change_materials.zip
|
|
194
|
Mon Feb 13 14:56:57 2023 |
Jack Tillman | Building - Physics Plots for 9_50, 13_84, 18_89, 19_96, 29_87 Antennae | Physics plots were created for 300K events from the higher statistic simulation results for the 9_50, 13_84, 18_89, 19_96, and 29_87 antennae.
The simulation was inaccurate because a discrepancy exists between the higher simulation physics plots and the physics plots created for the 300K event results currently in the GENETIS Loop. This can be seen in the attached pdf file.
The simulation may have been inaccurate due to incorrect gain files.
The higher statistic simulations must be rerun for the 9_50, 13_84, 18_89, 19_96, and 29_87 antennae.
|
| Attachment 1: Genetis_Physics_Plot_Comparison_Slides.pdf
|
|
230
|
Thu Jul 6 14:52:09 2023 |
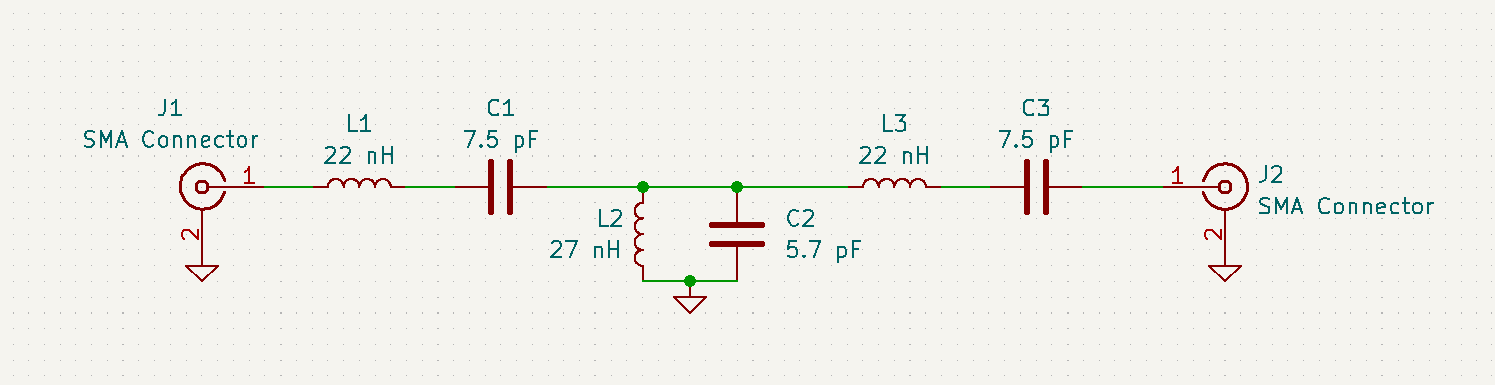
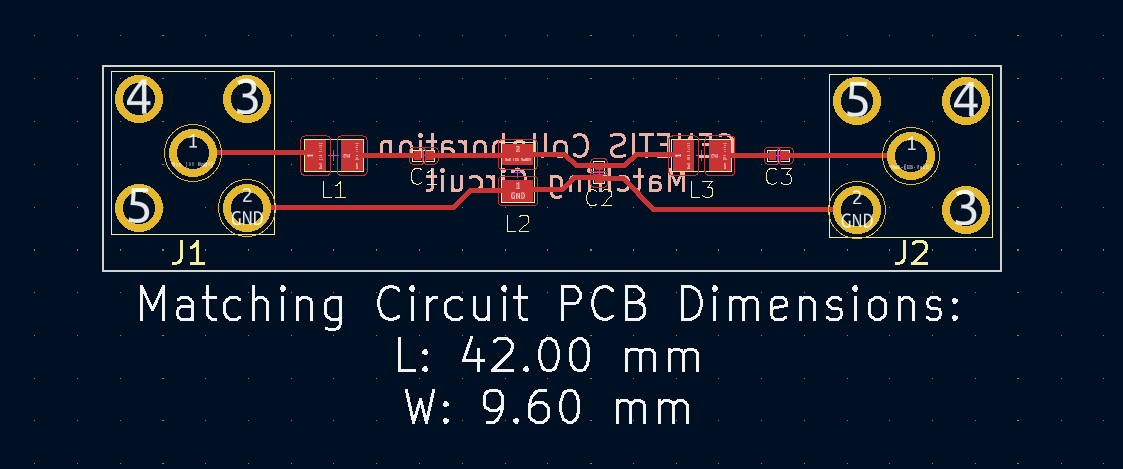
Jack Tillman | Building - Matching Circuit Schematic, PCB, and components | Attached are images of the matching circuit schematic and PCB design. A parts list is also attached in .pdf and .csv format. The .csv format can be imported into Digikey if necessary.
Table of component values:
| Inductors |
Capacitors |
SMA Connectors |
| 22 nH |
7.5 pF |
50 Ω |
| 27 nH |
5.7 pF |
50 Ω |
|
| Attachment 1: Matching_Circuit_Schematic.png
|  |
| Attachment 2: Matching_Circuit_PCB.png
|  |
| Attachment 3: Matching_Circuit_DigiKey_Cart.pdf
|
| Attachment 4: Matching_Circuit_DigiKey_Cart.csv
|
Index,Quantity,Part Number,Manufacturer Part Number,Description,Available,Backorder,Unit Price,Extended Price USD
1,3,WM26450-ND,733910083,"SMA RA JACK, PCB",3,0,5.1,15.3
2,4,490-GJM0335C1E7R5BB01DCT-ND,GJM0335C1E7R5BB01D,CAP CER 7.5PF 25V C0G/NP0 0201,4,0,0.1,0.4
3,2,490-17198-1-ND,GJM0335C1H5R7CB01D,CAP CER 5.7PF 50V C0G/NP0 0201,2,0,0.09,0.18
4,2,490-16032-1-ND,LQW2BAN27NJ00L,FIXED IND 27NH 2A 70 MOHM SMD,2,0,0.42,0.84
5,4,490-16028-1-ND,LQW2BAN22NJ00L,FIXED IND 22NH 1.9A 70 MOHM SMD,4,0,0.42,1.68
|
|
193
|
Fri Feb 10 15:55:53 2023 |
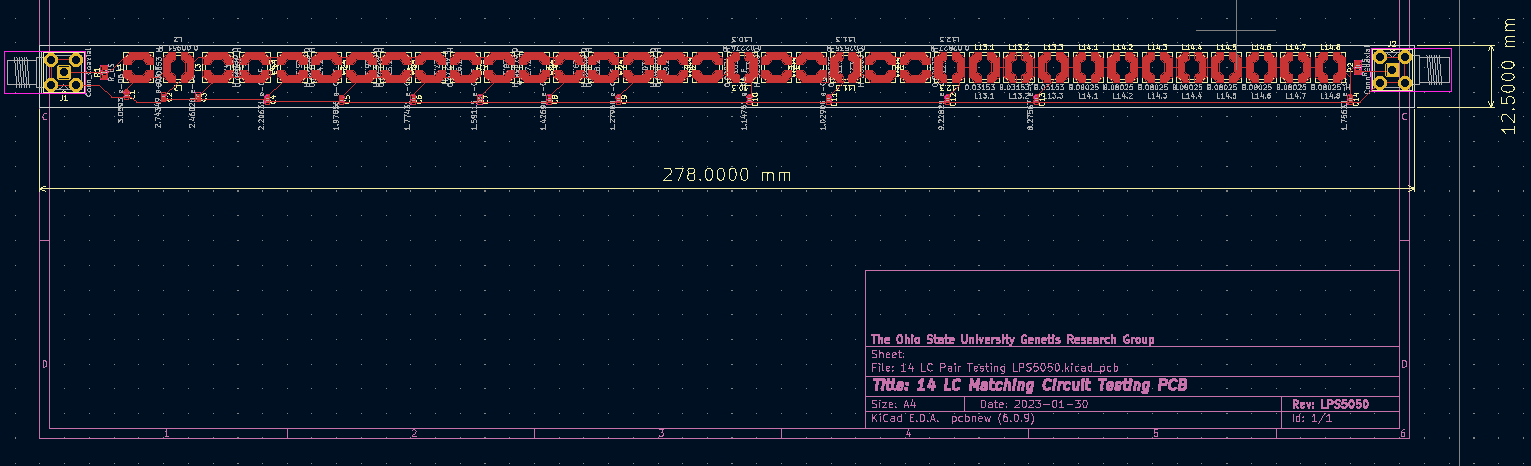
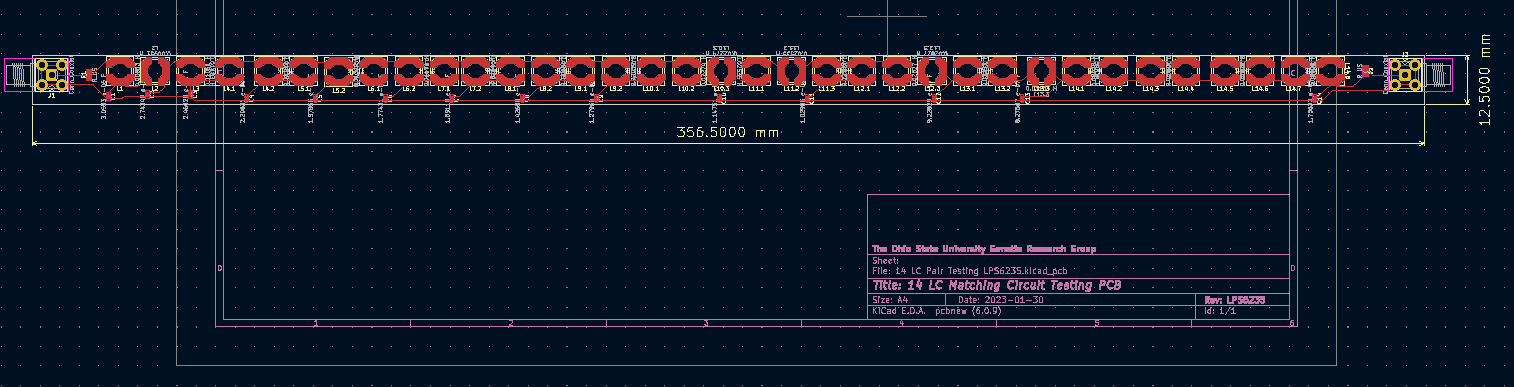
Jack Tillman | Building - Matching Circuit PCB | I've completed laying out the PCB for the 14-rung matching circuit. Attached are png files of two PCB designs. One uses LPS5050 inductors while the other uses LPS6235 inductors. The dimensions shown are in millimeters. |
| Attachment 1: LPS5050_Dimensioned_PCB.png
|  |
| Attachment 2: LPS6235_Dimensioned_PCB.png
|  |
|
189
|
Tue Jan 31 11:30:54 2023 |
Audrey Zinn | Building | Attached are the XF files for the top 5 individuals from the 2022_12_29 crazy sides run. In order from best to 5th best:
1: Gen 29 Indiv 87
2: Gen 19 Indiv 96
3: Gen 13 Indiv 84
4: Gen 9 Indiv 50
5: Gen 18 Indiv 89
These can also be found in /fs/ess/PAS1960/BiconeEvolutionOSC/BiconeEvolution/current_antenna_evo_build/XF_Loop/Evolutionary_Loop/Run_Outputs/2022_12_29/2022_12_29.xf in the corresponding directories. |
| Attachment 1: XF_model_13_84.txt
|
freq : 83.33 MHz
SWR : 1.965000
Theta Phi Gain(dB) Gain Phase(deg)
0 0 -64.90 0.00 -105.60
5 0 -19.63 0.01 -105.58
10 0 -13.64 0.04 -105.51
15 0 -10.17 0.10 -105.38
20 0 -7.75 0.17 -105.21
25 0 -5.90 0.26 -104.96
30 0 -4.43 0.36 -104.65
35 0 -3.23 0.48 -104.25
40 0 -2.22 0.60 -103.75
45 0 -1.38 0.73 -103.11
50 0 -0.67 0.86 -102.31
55 0 -0.08 0.98 -101.26
60 0 0.42 1.10 -99.87
65 0 0.82 1.21 -97.94
70 0 1.14 1.30 -95.09
75 0 1.39 1.38 -90.53
80 0 1.56 1.43 -82.17
85 0 1.67 1.47 -63.52
90 0 1.70 1.48 -19.53
95 0 1.67 1.47 24.56
100 0 1.56 1.43 43.26
105 0 1.39 1.38 51.64
110 0 1.14 1.30 56.22
115 0 0.82 1.21 59.07
120 0 0.41 1.10 61.01
125 0 -0.08 0.98 62.41
130 0 -0.68 0.86 63.46
135 0 -1.39 0.73 64.28
140 0 -2.23 0.60 64.92
145 0 -3.23 0.48 65.43
150 0 -4.44 0.36 65.84
155 0 -5.91 0.26 66.17
160 0 -7.76 0.17 66.42
165 0 -10.19 0.10 66.62
170 0 -13.66 0.04 66.75
175 0 -19.66 0.01 66.84
180 0 -64.87 0.00 66.87
0 5 -64.33 0.00 -107.33
5 5 -19.63 0.01 -107.32
10 5 -13.64 0.04 -107.27
15 5 -10.17 0.10 -107.18
20 5 -7.75 0.17 -107.05
25 5 -5.90 0.26 -106.88
30 5 -4.43 0.36 -106.65
35 5 -3.23 0.48 -106.37
40 5 -2.22 0.60 -106.01
45 5 -1.38 0.73 -105.57
50 5 -0.67 0.86 -105.00
55 5 -0.08 0.98 -104.28
60 5 0.42 1.10 -103.31
65 5 0.82 1.21 -101.97
70 5 1.14 1.30 -99.99
75 5 1.39 1.38 -96.77
80 5 1.56 1.43 -90.60
85 5 1.67 1.47 -74.77
90 5 1.70 1.48 -18.14
95 5 1.67 1.47 36.69
100 5 1.56 1.43 51.96
105 5 1.39 1.38 57.99
110 5 1.14 1.30 61.17
115 5 0.82 1.21 63.13
120 5 0.41 1.10 64.46
125 5 -0.08 0.98 65.43
130 5 -0.68 0.86 66.16
135 5 -1.39 0.73 66.73
140 5 -2.23 0.60 67.19
145 5 -3.23 0.48 67.56
150 5 -4.44 0.36 67.85
155 5 -5.91 0.26 68.09
160 5 -7.76 0.17 68.27
165 5 -10.19 0.10 68.42
170 5 -13.66 0.04 68.52
175 5 -19.66 0.01 68.58
180 5 -64.30 0.00 68.61
0 10 -63.85 0.00 -109.51
5 10 -19.63 0.01 -109.51
10 10 -13.64 0.04 -109.49
15 10 -10.17 0.10 -109.45
20 10 -7.75 0.17 -109.38
25 10 -5.90 0.26 -109.30
30 10 -4.43 0.36 -109.19
35 10 -3.23 0.48 -109.06
40 10 -2.22 0.60 -108.89
45 10 -1.38 0.73 -108.69
50 10 -0.67 0.86 -108.44
55 10 -0.08 0.98 -108.13
60 10 0.42 1.10 -107.73
65 10 0.82 1.21 -107.19
70 10 1.14 1.30 -106.41
75 10 1.39 1.38 -105.17
80 10 1.56 1.43 -102.75
85 10 1.67 1.47 -95.68
90 10 1.70 1.48 -10.72
95 10 1.67 1.47 57.68
100 10 1.56 1.43 64.05
105 10 1.39 1.38 66.34
110 10 1.14 1.30 67.55
115 10 0.82 1.21 68.31
120 10 0.41 1.10 68.85
125 10 -0.08 0.98 69.26
130 10 -0.68 0.86 69.58
135 10 -1.39 0.73 69.84
140 10 -2.23 0.60 70.06
145 10 -3.23 0.48 70.24
150 10 -4.44 0.36 70.39
155 10 -5.91 0.26 70.51
160 10 -7.76 0.17 70.61
165 10 -10.19 0.10 70.69
170 10 -13.66 0.04 70.75
175 10 -19.66 0.01 70.78
180 10 -63.82 0.00 70.80
0 15 -63.45 0.00 -112.40
5 15 -19.63 0.01 -112.41
10 15 -13.64 0.04 -112.42
15 15 -10.17 0.10 -112.44
20 15 -7.75 0.17 -112.47
25 15 -5.90 0.26 -112.50
30 15 -4.43 0.36 -112.55
35 15 -3.23 0.48 -112.61
40 15 -2.22 0.60 -112.70
45 15 -1.38 0.73 -112.83
50 15 -0.67 0.86 -113.01
55 15 -0.08 0.98 -113.26
60 15 0.42 1.10 -113.63
65 15 0.82 1.21 -114.19
70 15 1.14 1.30 -115.09
75 15 1.39 1.38 -116.67
80 15 1.56 1.43 -119.98
85 15 1.67 1.47 -130.06
90 15 1.70 1.48 150.28
95 15 1.67 1.47 88.84
100 15 1.56 1.43 80.40
105 15 1.39 1.38 77.44
110 15 1.14 1.30 75.99
115 15 0.82 1.21 75.16
120 15 0.41 1.10 74.65
125 15 -0.08 0.98 74.32
130 15 -0.68 0.86 74.09
135 15 -1.39 0.73 73.95
140 15 -2.23 0.60 73.85
145 15 -3.23 0.48 73.78
150 15 -4.44 0.36 73.74
155 15 -5.91 0.26 73.71
160 15 -7.76 0.17 73.70
165 15 -10.19 0.10 73.69
170 15 -13.66 0.04 73.69
175 15 -19.66 0.01 73.70
180 15 -63.42 0.00 73.71
0 20 -63.14 0.00 -116.43
5 20 -19.63 0.01 -116.46
10 20 -13.64 0.04 -116.53
15 20 -10.17 0.10 -116.63
20 20 -7.75 0.17 -116.78
25 20 -5.90 0.26 -116.98
30 20 -4.43 0.36 -117.25
35 20 -3.23 0.48 -117.59
40 20 -2.22 0.60 -118.04
45 20 -1.38 0.73 -118.62
50 20 -0.67 0.86 -119.39
55 20 -0.08 0.98 -120.42
60 20 0.42 1.10 -121.84
65 20 0.82 1.21 -123.88
70 20 1.14 1.30 -126.97
75 20 1.39 1.38 -132.07
80 20 1.56 1.43 -141.54
85 20 1.67 1.47 -162.14
90 20 1.70 1.48 155.74
95 20 1.67 1.47 117.62
100 20 1.56 1.43 100.14
105 20 1.39 1.38 91.86
110 20 1.14 1.30 87.29
115 20 0.82 1.21 84.47
120 20 0.41 1.10 82.60
125 20 -0.08 0.98 81.29
130 20 -0.68 0.86 80.35
135 20 -1.39 0.73 79.64
140 20 -2.23 0.60 79.12
145 20 -3.23 0.48 78.72
150 20 -4.44 0.36 78.41
155 20 -5.91 0.26 78.18
160 20 -7.76 0.17 78.02
165 20 -10.19 0.10 77.90
170 20 -13.66 0.04 77.82
175 20 -19.66 0.01 77.78
180 20 -63.11 0.00 77.78
0 25 -62.89 0.00 -122.51
5 25 -19.63 0.01 -122.57
10 25 -13.64 0.04 -122.71
15 25 -10.17 0.10 -122.95
20 25 -7.75 0.17 -123.28
25 25 -5.90 0.26 -123.72
30 25 -4.43 0.36 -124.31
35 25 -3.23 0.48 -125.05
40 25 -2.22 0.60 -126.00
45 25 -1.38 0.73 -127.22
50 25 -0.67 0.86 -128.80
55 25 -0.08 0.98 -130.86
60 25 0.42 1.10 -133.62
65 25 0.82 1.21 -137.41
70 25 1.14 1.30 -142.79
75 25 1.39 1.38 -150.72
80 25 1.56 1.43 -162.72
85 25 1.67 1.47 179.55
90 25 1.70 1.48 157.02
95 25 1.67 1.47 135.52
100 25 1.56 1.43 119.63
105 25 1.39 1.38 109.06
110 25 1.14 1.30 102.05
115 25 0.82 1.21 97.25
120 25 0.41 1.10 93.83
125 25 -0.08 0.98 91.34
130 25 -0.68 0.86 89.46
135 25 -1.39 0.73 88.03
140 25 -2.23 0.60 86.93
145 25 -3.23 0.48 86.07
150 25 -4.44 0.36 85.40
155 25 -5.91 0.26 84.89
160 25 -7.76 0.17 84.50
165 25 -10.19 0.10 84.22
170 25 -13.66 0.04 84.04
175 25 -19.66 0.01 83.94
180 25 -62.87 0.00 83.92
0 30 -62.72 0.00 -132.56
5 30 -19.63 0.01 -132.68
10 30 -13.64 0.04 -132.94
15 30 -10.17 0.10 -133.37
20 30 -7.75 0.17 -133.97
25 30 -5.90 0.26 -134.77
30 30 -4.43 0.36 -135.79
35 30 -3.23 0.48 -137.09
40 30 -2.22 0.60 -138.71
45 30 -1.38 0.73 -140.72
50 30 -0.67 0.86 -143.24
55 30 -0.08 0.98 -146.39
60 30 0.42 1.10 -150.34
65 30 0.82 1.21 -155.35
70 30 1.14 1.30 -161.66
75 30 1.39 1.38 -169.57
80 30 1.56 1.43 -179.20
85 30 1.67 1.47 169.63
90 30 1.70 1.48 157.66
95 30 1.67 1.47 145.96
100 30 1.56 1.43 135.50
105 30 1.39 1.38 126.73
110 30 1.14 1.30 119.66
115 30 0.82 1.21 114.04
120 30 0.41 1.10 109.61
125 30 -0.08 0.98 106.09
130 30 -0.68 0.86 103.30
135 30 -1.39 0.73 101.07
140 30 -2.23 0.60 99.29
145 30 -3.23 0.48 97.86
150 30 -4.44 0.36 96.72
155 30 -5.91 0.26 95.83
160 30 -7.76 0.17 95.15
165 30 -10.19 0.10 94.66
170 30 -13.66 0.04 94.33
175 30 -19.66 0.01 94.15
180 30 -62.70 0.00 94.13
0 35 -62.62 0.00 -150.86
5 35 -19.63 0.01 -151.07
10 35 -13.64 0.04 -151.49
15 35 -10.17 0.10 -152.14
20 35 -7.75 0.17 -153.03
25 35 -5.90 0.26 -154.18
30 35 -4.43 0.36 -155.60
35 35 -3.23 0.48 -157.32
40 35 -2.22 0.60 -159.38
45 35 -1.38 0.73 -161.80
50 35 -0.67 0.86 -164.62
55 35 -0.08 0.98 -167.87
60 35 0.42 1.10 -171.57
65 35 0.82 1.21 -175.73
70 35 1.14 1.30 179.66
75 35 1.39 1.38 174.64
80 35 1.56 1.43 169.28
85 35 1.67 1.47 163.72
90 35 1.70 1.48 158.08
95 35 1.67 1.47 152.52
100 35 1.56 1.43 147.18
105 35 1.39 1.38 142.18
110 35 1.14 1.30 137.59
115 35 0.82 1.21 133.45
120 35 0.41 1.10 129.78
125 35 -0.08 0.98 126.55
130 35 -0.68 0.86 123.75
135 35 -1.39 0.73 121.34
140 35 -2.23 0.60 119.29
145 35 -3.23 0.48 117.57
150 35 -4.44 0.36 116.14
155 35 -5.91 0.26 114.99
160 35 -7.76 0.17 114.09
165 35 -10.19 0.10 113.43
170 35 -13.66 0.04 113.00
175 35 -19.66 0.01 112.78
180 35 -62.60 0.00 112.77
0 40 -62.58 0.00 177.48
... 159721 more lines ...
|
| Attachment 2: XF_model_9_50.txt
|
freq : 83.33 MHz
SWR : 1.965000
Theta Phi Gain(dB) Gain Phase(deg)
0 0 -65.66 0.00 68.97
5 0 -23.00 0.01 68.99
10 0 -17.02 0.02 69.05
15 0 -13.55 0.04 69.15
20 0 -11.13 0.08 69.30
25 0 -9.29 0.12 69.49
30 0 -7.82 0.17 69.75
35 0 -6.63 0.22 70.06
40 0 -5.63 0.27 70.43
45 0 -4.79 0.33 70.90
50 0 -4.09 0.39 71.47
55 0 -3.50 0.45 72.18
60 0 -3.01 0.50 73.09
65 0 -2.61 0.55 74.31
70 0 -2.29 0.59 76.06
75 0 -2.05 0.62 78.83
80 0 -1.88 0.65 84.06
85 0 -1.78 0.66 97.84
90 0 -1.74 0.67 159.48
95 0 -1.78 0.66 -139.66
100 0 -1.88 0.65 -126.04
105 0 -2.05 0.62 -120.85
110 0 -2.30 0.59 -118.08
115 0 -2.61 0.55 -116.34
120 0 -3.02 0.50 -115.12
125 0 -3.51 0.45 -114.20
130 0 -4.10 0.39 -113.48
135 0 -4.80 0.33 -112.91
140 0 -5.63 0.27 -112.44
145 0 -6.63 0.22 -112.05
150 0 -7.83 0.16 -111.73
155 0 -9.30 0.12 -111.47
160 0 -11.14 0.08 -111.26
165 0 -13.57 0.04 -111.10
170 0 -17.04 0.02 -110.98
175 0 -23.04 0.00 -110.91
180 0 -65.62 0.00 -110.88
0 5 -66.34 0.00 70.70
5 5 -23.00 0.01 70.72
10 5 -17.02 0.02 70.80
15 5 -13.55 0.04 70.93
20 5 -11.13 0.08 71.12
25 5 -9.29 0.12 71.39
30 5 -7.82 0.17 71.72
35 5 -6.63 0.22 72.14
40 5 -5.63 0.27 72.65
45 5 -4.79 0.33 73.29
50 5 -4.09 0.39 74.09
55 5 -3.50 0.45 75.11
60 5 -3.01 0.50 76.44
65 5 -2.61 0.55 78.24
70 5 -2.29 0.59 80.87
75 5 -2.05 0.62 85.05
80 5 -1.88 0.65 92.78
85 5 -1.78 0.66 110.94
90 5 -1.74 0.67 160.03
95 5 -1.78 0.66 -152.08
100 5 -1.88 0.65 -134.51
105 5 -2.05 0.62 -126.96
110 5 -2.30 0.59 -122.85
115 5 -2.61 0.55 -120.26
120 5 -3.02 0.50 -118.46
125 5 -3.51 0.45 -117.14
130 5 -4.10 0.39 -116.13
135 5 -4.80 0.33 -115.32
140 5 -5.63 0.27 -114.67
145 5 -6.63 0.22 -114.15
150 5 -7.83 0.16 -113.72
155 5 -9.30 0.12 -113.38
160 5 -11.14 0.08 -113.10
165 5 -13.57 0.04 -112.89
170 5 -17.04 0.02 -112.75
175 5 -23.04 0.00 -112.66
180 5 -66.29 0.00 -112.62
0 10 -67.14 0.00 72.17
5 10 -23.00 0.01 72.19
10 10 -17.02 0.02 72.28
15 10 -13.55 0.04 72.45
20 10 -11.13 0.08 72.68
25 10 -9.29 0.12 72.99
30 10 -7.82 0.17 73.40
35 10 -6.63 0.22 73.91
40 10 -5.63 0.27 74.54
45 10 -4.79 0.33 75.33
50 10 -4.09 0.39 76.32
55 10 -3.50 0.45 77.59
60 10 -3.01 0.50 79.26
65 10 -2.61 0.55 81.54
70 10 -2.29 0.59 84.86
75 10 -2.05 0.62 90.10
80 10 -1.88 0.65 99.46
85 10 -1.78 0.66 119.21
90 10 -1.74 0.67 160.25
95 10 -1.78 0.66 -159.82
100 10 -1.88 0.65 -140.91
105 10 -2.05 0.62 -131.87
110 10 -2.30 0.59 -126.77
115 10 -2.61 0.55 -123.52
120 10 -3.02 0.50 -121.27
125 10 -3.51 0.45 -119.61
130 10 -4.10 0.39 -118.35
135 10 -4.80 0.33 -117.36
140 10 -5.63 0.27 -116.57
145 10 -6.63 0.22 -115.93
150 10 -7.83 0.16 -115.41
155 10 -9.30 0.12 -114.99
160 10 -11.14 0.08 -114.67
165 10 -13.57 0.04 -114.42
170 10 -17.04 0.02 -114.25
175 10 -23.04 0.00 -114.14
180 10 -67.09 0.00 -114.10
0 15 -68.08 0.00 73.45
5 15 -23.00 0.01 73.48
10 15 -17.02 0.02 73.58
15 15 -13.55 0.04 73.77
20 15 -11.13 0.08 74.04
25 15 -9.29 0.12 74.40
30 15 -7.82 0.17 74.86
35 15 -6.63 0.22 75.45
40 15 -5.63 0.27 76.18
45 15 -4.79 0.33 77.10
50 15 -4.09 0.39 78.26
55 15 -3.50 0.45 79.74
60 15 -3.01 0.50 81.70
65 15 -2.61 0.55 84.37
70 15 -2.29 0.59 88.24
75 15 -2.05 0.62 94.27
80 15 -1.88 0.65 104.69
85 15 -1.78 0.66 124.77
90 15 -1.74 0.67 160.35
95 15 -1.78 0.66 -165.00
100 15 -1.88 0.65 -145.87
105 15 -2.05 0.62 -135.89
110 15 -2.30 0.59 -130.06
115 15 -2.61 0.55 -126.29
120 15 -3.02 0.50 -123.67
125 15 -3.51 0.45 -121.74
130 15 -4.10 0.39 -120.28
135 15 -4.80 0.33 -119.13
140 15 -5.63 0.27 -118.21
145 15 -6.63 0.22 -117.47
150 15 -7.83 0.16 -116.88
155 15 -9.30 0.12 -116.40
160 15 -11.14 0.08 -116.03
165 15 -13.57 0.04 -115.75
170 15 -17.04 0.02 -115.55
175 15 -23.04 0.00 -115.43
180 15 -68.02 0.00 -115.39
0 20 -69.19 0.00 74.59
5 20 -23.00 0.01 74.62
10 20 -17.02 0.02 74.74
15 20 -13.55 0.04 74.95
20 20 -11.13 0.08 75.25
25 20 -9.29 0.12 75.65
30 20 -7.82 0.17 76.17
35 20 -6.63 0.22 76.82
40 20 -5.63 0.27 77.64
45 20 -4.79 0.33 78.67
50 20 -4.09 0.39 79.97
55 20 -3.50 0.45 81.64
60 20 -3.01 0.50 83.84
65 20 -2.61 0.55 86.85
70 20 -2.29 0.59 91.16
75 20 -2.05 0.62 97.79
80 20 -1.88 0.65 108.88
85 20 -1.78 0.66 128.73
90 20 -1.74 0.67 160.41
95 20 -1.78 0.66 -168.71
100 20 -1.88 0.65 -149.83
105 20 -2.05 0.62 -139.26
110 20 -2.30 0.59 -132.89
115 20 -2.61 0.55 -128.71
120 20 -3.02 0.50 -125.78
125 20 -3.51 0.45 -123.62
130 20 -4.10 0.39 -121.98
135 20 -4.80 0.33 -120.69
140 20 -5.63 0.27 -119.66
145 20 -6.63 0.22 -118.84
150 20 -7.83 0.16 -118.18
155 20 -9.30 0.12 -117.66
160 20 -11.14 0.08 -117.24
165 20 -13.57 0.04 -116.93
170 20 -17.04 0.02 -116.72
175 20 -23.04 0.00 -116.58
180 20 -69.13 0.00 -116.53
0 25 -70.51 0.00 75.63
5 25 -23.00 0.01 75.67
10 25 -17.02 0.02 75.80
15 25 -13.56 0.04 76.02
20 25 -11.13 0.08 76.35
25 25 -9.29 0.12 76.79
30 25 -7.82 0.17 77.35
35 25 -6.63 0.22 78.07
40 25 -5.63 0.27 78.97
45 25 -4.79 0.33 80.10
50 25 -4.09 0.39 81.52
55 25 -3.50 0.45 83.36
60 25 -3.01 0.50 85.77
65 25 -2.61 0.55 89.05
70 25 -2.29 0.59 93.74
75 25 -2.05 0.62 100.82
80 25 -1.88 0.65 112.32
85 25 -1.78 0.66 131.71
90 25 -1.74 0.67 160.43
95 25 -1.78 0.66 -171.53
100 25 -1.88 0.65 -153.09
105 25 -2.05 0.62 -142.16
110 25 -2.30 0.59 -135.37
115 25 -2.61 0.55 -130.85
120 25 -3.02 0.50 -127.66
125 25 -3.51 0.45 -125.31
130 25 -4.10 0.39 -123.51
135 25 -4.80 0.33 -122.10
140 25 -5.63 0.27 -120.98
145 25 -6.63 0.22 -120.09
150 25 -7.83 0.16 -119.37
155 25 -9.30 0.12 -118.79
160 25 -11.14 0.08 -118.35
165 25 -13.57 0.04 -118.01
170 25 -17.04 0.02 -117.77
175 25 -23.04 0.00 -117.63
180 25 -70.44 0.00 -117.58
0 30 -72.08 0.00 76.59
5 30 -23.00 0.01 76.63
10 30 -17.02 0.02 76.77
15 30 -13.56 0.04 77.02
20 30 -11.13 0.08 77.37
25 30 -9.29 0.12 77.84
30 30 -7.82 0.17 78.45
35 30 -6.63 0.22 79.23
40 30 -5.63 0.27 80.20
45 30 -4.79 0.33 81.41
50 30 -4.09 0.39 82.96
55 30 -3.50 0.45 84.93
60 30 -3.01 0.50 87.53
65 30 -2.61 0.55 91.06
70 30 -2.29 0.59 96.04
75 30 -2.05 0.62 103.48
80 30 -1.88 0.65 115.21
85 30 -1.78 0.66 134.04
90 30 -1.74 0.67 160.43
95 30 -1.78 0.66 -173.76
100 30 -1.88 0.65 -155.84
105 30 -2.05 0.62 -144.70
110 30 -2.30 0.59 -137.59
115 30 -2.61 0.55 -132.79
120 30 -3.02 0.50 -129.38
125 30 -3.51 0.45 -126.85
130 30 -4.10 0.39 -124.91
135 30 -4.80 0.33 -123.40
140 30 -5.63 0.27 -122.20
145 30 -6.63 0.22 -121.23
150 30 -7.83 0.16 -120.46
155 30 -9.30 0.12 -119.84
160 30 -11.14 0.08 -119.36
165 30 -13.57 0.04 -119.00
170 30 -17.04 0.02 -118.75
175 30 -23.04 0.00 -118.60
180 30 -72.01 0.00 -118.54
0 35 -73.95 0.00 77.50
5 35 -23.00 0.01 77.55
10 35 -17.02 0.02 77.70
15 35 -13.56 0.04 77.95
20 35 -11.13 0.08 78.33
25 35 -9.29 0.12 78.83
30 35 -7.82 0.17 79.49
35 35 -6.63 0.22 80.31
40 35 -5.63 0.27 81.35
45 35 -4.79 0.33 82.65
50 35 -4.09 0.39 84.29
55 35 -3.50 0.45 86.40
60 35 -3.01 0.50 89.17
65 35 -2.61 0.55 92.90
70 35 -2.29 0.59 98.15
75 35 -2.05 0.62 105.86
80 35 -1.88 0.65 117.70
85 35 -1.78 0.66 135.93
90 35 -1.74 0.67 160.42
95 35 -1.78 0.66 -175.60
100 35 -1.88 0.65 -158.22
105 35 -2.05 0.62 -146.97
110 35 -2.30 0.59 -139.62
115 35 -2.61 0.55 -134.58
120 35 -3.02 0.50 -130.97
125 35 -3.51 0.45 -128.29
130 35 -4.10 0.39 -126.23
135 35 -4.80 0.33 -124.62
140 35 -5.63 0.27 -123.33
145 35 -6.63 0.22 -122.31
150 35 -7.83 0.16 -121.49
155 35 -9.30 0.12 -120.83
160 35 -11.14 0.08 -120.32
165 35 -13.57 0.04 -119.94
170 35 -17.04 0.02 -119.67
175 35 -23.04 0.00 -119.51
180 35 -73.87 0.00 -119.46
0 40 -76.10 0.00 78.37
... 159721 more lines ...
|
| Attachment 3: XF_model_18_89.txt
|
freq : 83.33 MHz
SWR : 1.965000
Theta Phi Gain(dB) Gain Phase(deg)
0 0 -300.00 0.00 -25.63
5 0 -300.00 0.00 -25.69
10 0 -300.00 0.00 -25.72
15 0 -300.00 0.00 -25.73
20 0 -300.00 0.00 -25.71
25 0 -300.00 0.00 -25.66
30 0 -300.00 0.00 -25.58
35 0 -300.00 0.00 -25.48
40 0 -300.00 0.00 -25.35
45 0 -300.00 0.00 -25.20
50 0 -300.00 0.00 -25.02
55 0 -300.00 0.00 -24.82
60 0 -300.00 0.00 -24.59
65 0 -300.00 0.00 -24.33
70 0 -300.00 0.00 -24.06
75 0 -300.00 0.00 -23.76
80 0 -300.00 0.00 -23.44
85 0 -300.00 0.00 -23.11
90 0 -300.00 0.00 -22.76
95 0 -300.00 0.00 -22.39
100 0 -300.00 0.00 -22.02
105 0 -300.00 0.00 -21.63
110 0 -300.00 0.00 -21.24
115 0 -300.00 0.00 -20.85
120 0 -300.00 0.00 -20.46
125 0 -300.00 0.00 -20.07
130 0 -300.00 0.00 -19.70
135 0 -300.00 0.00 -19.34
140 0 -300.00 0.00 -18.99
145 0 -300.00 0.00 -18.67
150 0 -300.00 0.00 -18.37
155 0 -300.00 0.00 -18.09
160 0 -300.00 0.00 -17.85
165 0 -300.00 0.00 -17.64
170 0 -300.00 0.00 -17.47
175 0 -300.00 0.00 -17.33
180 0 -300.00 0.00 -17.23
0 5 -300.00 0.00 -57.01
5 5 -300.00 0.00 -56.84
10 5 -300.00 0.00 -56.42
15 5 -300.00 0.00 -55.73
20 5 -300.00 0.00 -54.77
25 5 -300.00 0.00 -53.53
30 5 -300.00 0.00 -52.01
35 5 -300.00 0.00 -50.20
40 5 -300.00 0.00 -48.07
45 5 -300.00 0.00 -45.62
50 5 -300.00 0.00 -42.84
55 5 -300.00 0.00 -39.72
60 5 -300.00 0.00 -36.29
65 5 -300.00 0.00 -32.55
70 5 -300.00 0.00 -28.55
75 5 -300.00 0.00 -24.35
80 5 -300.00 0.00 -20.01
85 5 -300.00 0.00 -15.63
90 5 -300.00 0.00 -11.28
95 5 -300.00 0.00 -7.06
100 5 -300.00 0.00 -3.05
105 5 -300.00 0.00 0.71
110 5 -300.00 0.00 4.17
115 5 -300.00 0.00 7.31
120 5 -300.00 0.00 10.11
125 5 -300.00 0.00 12.60
130 5 -300.00 0.00 14.77
135 5 -300.00 0.00 16.65
140 5 -300.00 0.00 18.25
145 5 -300.00 0.00 19.59
150 5 -300.00 0.00 20.70
155 5 -300.00 0.00 21.58
160 5 -300.00 0.00 22.25
165 5 -300.00 0.00 22.74
170 5 -300.00 0.00 23.03
175 5 -300.00 0.00 23.15
180 5 -300.00 0.00 23.10
0 10 -300.00 0.00 -64.53
5 10 -300.00 0.00 -64.32
10 10 -300.00 0.00 -63.83
15 10 -300.00 0.00 -63.05
20 10 -300.00 0.00 -61.97
25 10 -300.00 0.00 -60.57
30 10 -300.00 0.00 -58.84
35 10 -300.00 0.00 -56.75
40 10 -300.00 0.00 -54.27
45 10 -300.00 0.00 -51.37
50 10 -300.00 0.00 -48.02
55 10 -300.00 0.00 -44.20
60 10 -300.00 0.00 -39.91
65 10 -300.00 0.00 -35.16
70 10 -300.00 0.00 -30.01
75 10 -300.00 0.00 -24.54
80 10 -300.00 0.00 -18.89
85 10 -300.00 0.00 -13.22
90 10 -300.00 0.00 -7.66
95 10 -300.00 0.00 -2.37
100 10 -300.00 0.00 2.54
105 10 -300.00 0.00 7.02
110 10 -300.00 0.00 11.02
115 10 -300.00 0.00 14.56
120 10 -300.00 0.00 17.64
125 10 -300.00 0.00 20.29
130 10 -300.00 0.00 22.57
135 10 -300.00 0.00 24.49
140 10 -300.00 0.00 26.11
145 10 -300.00 0.00 27.44
150 10 -300.00 0.00 28.52
155 10 -300.00 0.00 29.37
160 10 -300.00 0.00 30.01
165 10 -300.00 0.00 30.45
170 10 -300.00 0.00 30.71
175 10 -300.00 0.00 30.79
180 10 -300.00 0.00 30.71
0 15 -300.00 0.00 -67.78
5 15 -300.00 0.00 -67.56
10 15 -300.00 0.00 -67.05
15 15 -300.00 0.00 -66.24
20 15 -300.00 0.00 -65.13
25 15 -300.00 0.00 -63.69
30 15 -300.00 0.00 -61.90
35 15 -300.00 0.00 -59.72
40 15 -300.00 0.00 -57.12
45 15 -300.00 0.00 -54.05
50 15 -300.00 0.00 -50.47
55 15 -300.00 0.00 -46.35
60 15 -300.00 0.00 -41.67
65 15 -300.00 0.00 -36.45
70 15 -300.00 0.00 -30.73
75 15 -300.00 0.00 -24.64
80 15 -300.00 0.00 -18.34
85 15 -300.00 0.00 -12.03
90 15 -300.00 0.00 -5.90
95 15 -300.00 0.00 -0.13
100 15 -300.00 0.00 5.17
105 15 -300.00 0.00 9.92
110 15 -300.00 0.00 14.11
115 15 -300.00 0.00 17.75
120 15 -300.00 0.00 20.89
125 15 -300.00 0.00 23.58
130 15 -300.00 0.00 25.85
135 15 -300.00 0.00 27.76
140 15 -300.00 0.00 29.35
145 15 -300.00 0.00 30.65
150 15 -300.00 0.00 31.70
155 15 -300.00 0.00 32.52
160 15 -300.00 0.00 33.14
165 15 -300.00 0.00 33.56
170 15 -300.00 0.00 33.80
175 15 -300.00 0.00 33.87
180 15 -300.00 0.00 33.78
0 20 -300.00 0.00 -69.61
5 20 -300.00 0.00 -69.38
10 20 -300.00 0.00 -68.87
15 20 -300.00 0.00 -68.05
20 20 -300.00 0.00 -66.92
25 20 -300.00 0.00 -65.47
30 20 -300.00 0.00 -63.65
35 20 -300.00 0.00 -61.43
40 20 -300.00 0.00 -58.77
45 20 -300.00 0.00 -55.61
50 20 -300.00 0.00 -51.92
55 20 -300.00 0.00 -47.63
60 20 -300.00 0.00 -42.73
65 20 -300.00 0.00 -37.23
70 20 -300.00 0.00 -31.18
75 20 -300.00 0.00 -24.71
80 20 -300.00 0.00 -18.02
85 20 -300.00 0.00 -11.32
90 20 -300.00 0.00 -4.86
95 20 -300.00 0.00 1.18
100 20 -300.00 0.00 6.68
105 20 -300.00 0.00 11.57
110 20 -300.00 0.00 15.84
115 20 -300.00 0.00 19.54
120 20 -300.00 0.00 22.70
125 20 -300.00 0.00 25.38
130 20 -300.00 0.00 27.64
135 20 -300.00 0.00 29.54
140 20 -300.00 0.00 31.11
145 20 -300.00 0.00 32.39
150 20 -300.00 0.00 33.42
155 20 -300.00 0.00 34.23
160 20 -300.00 0.00 34.83
165 20 -300.00 0.00 35.24
170 20 -300.00 0.00 35.47
175 20 -300.00 0.00 35.54
180 20 -300.00 0.00 35.45
0 25 -300.00 0.00 -70.79
5 25 -300.00 0.00 -70.56
10 25 -300.00 0.00 -70.04
15 25 -300.00 0.00 -69.23
20 25 -300.00 0.00 -68.10
25 25 -300.00 0.00 -66.63
30 25 -300.00 0.00 -64.80
35 25 -300.00 0.00 -62.56
40 25 -300.00 0.00 -59.86
45 25 -300.00 0.00 -56.66
50 25 -300.00 0.00 -52.88
55 25 -300.00 0.00 -48.49
60 25 -300.00 0.00 -43.45
65 25 -300.00 0.00 -37.76
70 25 -300.00 0.00 -31.49
75 25 -300.00 0.00 -24.77
80 25 -300.00 0.00 -17.81
85 25 -300.00 0.00 -10.86
90 25 -300.00 0.00 -4.18
95 25 -300.00 0.00 2.03
100 25 -300.00 0.00 7.65
105 25 -300.00 0.00 12.62
110 25 -300.00 0.00 16.95
115 25 -300.00 0.00 20.67
120 25 -300.00 0.00 23.83
125 25 -300.00 0.00 26.51
130 25 -300.00 0.00 28.77
135 25 -300.00 0.00 30.65
140 25 -300.00 0.00 32.21
145 25 -300.00 0.00 33.48
150 25 -300.00 0.00 34.50
155 25 -300.00 0.00 35.30
160 25 -300.00 0.00 35.89
165 25 -300.00 0.00 36.30
170 25 -300.00 0.00 36.53
175 25 -300.00 0.00 36.60
180 25 -300.00 0.00 36.51
0 30 -300.00 0.00 -71.63
5 30 -300.00 0.00 -71.40
10 30 -300.00 0.00 -70.88
15 30 -300.00 0.00 -70.07
20 30 -300.00 0.00 -68.94
25 30 -300.00 0.00 -67.47
30 30 -300.00 0.00 -65.63
35 30 -300.00 0.00 -63.37
40 30 -300.00 0.00 -60.65
45 30 -300.00 0.00 -57.41
50 30 -300.00 0.00 -53.59
55 30 -300.00 0.00 -49.13
60 30 -300.00 0.00 -43.98
65 30 -300.00 0.00 -38.16
70 30 -300.00 0.00 -31.72
75 30 -300.00 0.00 -24.82
80 30 -300.00 0.00 -17.67
85 30 -300.00 0.00 -10.55
90 30 -300.00 0.00 -3.71
95 30 -300.00 0.00 2.62
100 30 -300.00 0.00 8.33
105 30 -300.00 0.00 13.35
110 30 -300.00 0.00 17.70
115 30 -300.00 0.00 21.43
120 30 -300.00 0.00 24.61
125 30 -300.00 0.00 27.28
130 30 -300.00 0.00 29.53
135 30 -300.00 0.00 31.40
140 30 -300.00 0.00 32.96
145 30 -300.00 0.00 34.22
150 30 -300.00 0.00 35.24
155 30 -300.00 0.00 36.03
160 30 -300.00 0.00 36.62
165 30 -300.00 0.00 37.03
170 30 -300.00 0.00 37.27
175 30 -300.00 0.00 37.34
180 30 -300.00 0.00 37.25
0 35 -300.00 0.00 -72.26
5 35 -300.00 0.00 -72.04
10 35 -300.00 0.00 -71.52
15 35 -300.00 0.00 -70.71
20 35 -300.00 0.00 -69.58
25 35 -300.00 0.00 -68.11
30 35 -300.00 0.00 -66.26
35 35 -300.00 0.00 -64.00
40 35 -300.00 0.00 -61.27
45 35 -300.00 0.00 -58.00
50 35 -300.00 0.00 -54.14
55 35 -300.00 0.00 -49.62
60 35 -300.00 0.00 -44.40
65 35 -300.00 0.00 -38.47
70 35 -300.00 0.00 -31.91
75 35 -300.00 0.00 -24.87
80 35 -300.00 0.00 -17.58
85 35 -300.00 0.00 -10.32
90 35 -300.00 0.00 -3.38
95 35 -300.00 0.00 3.04
100 35 -300.00 0.00 8.81
105 35 -300.00 0.00 13.87
110 35 -300.00 0.00 18.24
115 35 -300.00 0.00 21.98
120 35 -300.00 0.00 25.15
125 35 -300.00 0.00 27.83
130 35 -300.00 0.00 30.07
135 35 -300.00 0.00 31.94
140 35 -300.00 0.00 33.49
145 35 -300.00 0.00 34.76
150 35 -300.00 0.00 35.77
155 35 -300.00 0.00 36.57
160 35 -300.00 0.00 37.16
165 35 -300.00 0.00 37.57
170 35 -300.00 0.00 37.81
175 35 -300.00 0.00 37.89
180 35 -300.00 0.00 37.81
0 40 -300.00 0.00 -72.77
... 159721 more lines ...
|
| Attachment 4: XF_model_19_96.txt
|
freq : 83.33 MHz
SWR : 1.965000
Theta Phi Gain(dB) Gain Phase(deg)
0 0 -83.71 0.00 -158.06
5 0 -17.86 0.02 -153.75
10 0 -11.87 0.06 -148.66
15 0 -8.40 0.14 -144.50
20 0 -5.97 0.25 -140.71
25 0 -4.13 0.39 -136.31
30 0 -2.66 0.54 -131.91
35 0 -1.45 0.72 -127.38
40 0 -0.45 0.90 -123.42
45 0 0.39 1.09 -119.24
50 0 1.09 1.29 -115.23
55 0 1.69 1.47 -111.53
60 0 2.18 1.65 -107.68
65 0 2.59 1.81 -104.22
70 0 2.91 1.95 -100.95
75 0 3.15 2.07 -97.77
80 0 3.33 2.15 -94.90
85 0 3.43 2.20 -92.40
90 0 3.46 2.22 -89.68
95 0 3.43 2.20 -87.43
100 0 3.33 2.15 -85.51
105 0 3.15 2.07 -84.01
110 0 2.91 1.95 -82.20
115 0 2.59 1.81 -80.97
120 0 2.18 1.65 -79.70
125 0 1.69 1.47 -78.83
130 0 1.09 1.29 -77.85
135 0 0.39 1.09 -77.23
140 0 -0.45 0.90 -77.60
145 0 -1.45 0.72 -77.60
150 0 -2.66 0.54 -78.04
155 0 -4.13 0.39 -78.59
160 0 -5.97 0.25 -79.94
165 0 -8.40 0.14 -81.34
170 0 -11.87 0.07 -84.60
175 0 -17.86 0.02 -89.05
180 0 -83.70 0.00 -95.44
0 5 -83.74 0.00 -20.42
5 5 -17.86 0.02 -21.56
10 5 -11.87 0.06 -22.65
15 5 -8.40 0.14 -23.78
20 5 -5.97 0.25 -24.81
25 5 -4.13 0.39 -25.80
30 5 -2.66 0.54 -26.78
35 5 -1.45 0.72 -27.69
40 5 -0.45 0.90 -28.54
45 5 0.39 1.09 -29.38
50 5 1.09 1.29 -30.15
55 5 1.69 1.47 -30.88
60 5 2.18 1.65 -31.59
65 5 2.59 1.81 -32.25
70 5 2.91 1.95 -32.87
75 5 3.15 2.07 -33.45
80 5 3.33 2.15 -33.97
85 5 3.43 2.20 -34.45
90 5 3.46 2.22 -34.85
95 5 3.43 2.20 -35.18
100 5 3.33 2.15 -35.40
105 5 3.15 2.07 -35.52
110 5 2.91 1.95 -35.51
115 5 2.59 1.81 -35.38
120 5 2.18 1.65 -35.10
125 5 1.69 1.47 -34.70
130 5 1.09 1.29 -34.17
135 5 0.39 1.09 -33.48
140 5 -0.45 0.90 -32.71
145 5 -1.45 0.72 -31.84
150 5 -2.66 0.54 -30.83
155 5 -4.13 0.39 -29.77
160 5 -5.97 0.25 -28.61
165 5 -8.40 0.14 -27.42
170 5 -11.87 0.07 -26.17
175 5 -17.86 0.02 -24.90
180 5 -83.74 0.00 -23.60
0 10 -83.84 0.00 -19.91
5 10 -17.86 0.02 -20.98
10 10 -11.87 0.06 -22.04
15 10 -8.40 0.14 -23.08
20 10 -5.97 0.25 -24.09
25 10 -4.13 0.39 -25.07
30 10 -2.66 0.54 -26.00
35 10 -1.45 0.72 -26.89
40 10 -0.45 0.90 -27.75
45 10 0.39 1.09 -28.55
50 10 1.09 1.29 -29.32
55 10 1.69 1.47 -30.06
60 10 2.18 1.65 -30.76
65 10 2.59 1.81 -31.42
70 10 2.91 1.95 -32.05
75 10 3.15 2.07 -32.64
80 10 3.33 2.15 -33.17
85 10 3.43 2.20 -33.66
90 10 3.46 2.22 -34.07
95 10 3.43 2.20 -34.41
100 10 3.33 2.15 -34.64
105 10 3.15 2.07 -34.77
110 10 2.91 1.95 -34.79
115 10 2.59 1.81 -34.68
120 10 2.18 1.65 -34.44
125 10 1.69 1.47 -34.06
130 10 1.09 1.29 -33.55
135 10 0.39 1.09 -32.91
140 10 -0.45 0.90 -32.16
145 10 -1.45 0.72 -31.30
150 10 -2.66 0.54 -30.34
155 10 -4.13 0.39 -29.31
160 10 -5.97 0.25 -28.20
165 10 -8.40 0.14 -27.05
170 10 -11.87 0.07 -25.86
175 10 -17.86 0.02 -24.64
180 10 -83.84 0.00 -23.37
0 15 -84.01 0.00 -19.75
5 15 -17.86 0.02 -20.78
10 15 -11.87 0.06 -21.80
15 15 -8.40 0.14 -22.82
20 15 -5.97 0.25 -23.80
25 15 -4.13 0.39 -24.75
30 15 -2.66 0.54 -25.67
35 15 -1.45 0.72 -26.54
40 15 -0.45 0.90 -27.38
45 15 0.39 1.09 -28.17
50 15 1.09 1.29 -28.93
55 15 1.69 1.47 -29.66
60 15 2.18 1.65 -30.36
65 15 2.59 1.81 -31.01
70 15 2.91 1.95 -31.63
75 15 3.15 2.07 -32.21
80 15 3.33 2.15 -32.75
85 15 3.43 2.20 -33.22
90 15 3.46 2.22 -33.63
95 15 3.43 2.20 -33.96
100 15 3.33 2.15 -34.19
105 15 3.15 2.07 -34.33
110 15 2.91 1.95 -34.35
115 15 2.59 1.81 -34.24
120 15 2.18 1.65 -34.00
125 15 1.69 1.47 -33.65
130 15 1.09 1.29 -33.16
135 15 0.39 1.09 -32.54
140 15 -0.45 0.90 -31.82
145 15 -1.45 0.72 -30.99
150 15 -2.66 0.54 -30.07
155 15 -4.13 0.39 -29.06
160 15 -5.97 0.25 -28.00
165 15 -8.40 0.14 -26.87
170 15 -11.87 0.07 -25.71
175 15 -17.86 0.02 -24.52
180 15 -84.01 0.00 -23.30
0 20 -84.25 0.00 -19.66
5 20 -17.86 0.02 -20.66
10 20 -11.87 0.06 -21.65
15 20 -8.40 0.14 -22.63
20 20 -5.97 0.25 -23.59
25 20 -4.13 0.39 -24.52
30 20 -2.66 0.54 -25.42
35 20 -1.45 0.72 -26.27
40 20 -0.45 0.90 -27.10
45 20 0.39 1.09 -27.88
50 20 1.09 1.29 -28.63
55 20 1.69 1.47 -29.35
60 20 2.18 1.65 -30.03
65 20 2.59 1.81 -30.67
70 20 2.91 1.95 -31.28
75 20 3.15 2.07 -31.84
80 20 3.33 2.15 -32.36
85 20 3.43 2.20 -32.82
90 20 3.46 2.22 -33.21
95 20 3.43 2.20 -33.52
100 20 3.33 2.15 -33.75
105 20 3.15 2.07 -33.88
110 20 2.91 1.95 -33.89
115 20 2.59 1.81 -33.79
120 20 2.18 1.65 -33.57
125 20 1.69 1.47 -33.22
130 20 1.09 1.29 -32.76
135 20 0.39 1.09 -32.17
140 20 -0.45 0.90 -31.48
145 20 -1.45 0.72 -30.68
150 20 -2.66 0.54 -29.80
155 20 -4.13 0.39 -28.83
160 20 -5.97 0.25 -27.80
165 20 -8.40 0.14 -26.72
170 20 -11.87 0.07 -25.60
175 20 -17.86 0.02 -24.44
180 20 -84.25 0.00 -23.27
0 25 -84.56 0.00 -19.61
5 25 -17.86 0.02 -20.56
10 25 -11.87 0.06 -21.52
15 25 -8.40 0.14 -22.47
20 25 -5.97 0.25 -23.40
25 25 -4.13 0.39 -24.30
30 25 -2.66 0.54 -25.17
35 25 -1.45 0.72 -26.01
40 25 -0.45 0.90 -26.81
45 25 0.39 1.09 -27.58
50 25 1.09 1.29 -28.32
55 25 1.69 1.47 -29.01
60 25 2.18 1.65 -29.68
65 25 2.59 1.81 -30.31
70 25 2.91 1.95 -30.89
75 25 3.15 2.07 -31.44
80 25 3.33 2.15 -31.93
85 25 3.43 2.20 -32.37
90 25 3.46 2.22 -32.75
95 25 3.43 2.20 -33.04
100 25 3.33 2.15 -33.25
105 25 3.15 2.07 -33.37
110 25 2.91 1.95 -33.38
115 25 2.59 1.81 -33.28
120 25 2.18 1.65 -33.07
125 25 1.69 1.47 -32.74
130 25 1.09 1.29 -32.30
135 25 0.39 1.09 -31.75
140 25 -0.45 0.90 -31.09
145 25 -1.45 0.72 -30.35
150 25 -2.66 0.54 -29.50
155 25 -4.13 0.39 -28.58
160 25 -5.97 0.25 -27.60
165 25 -8.40 0.14 -26.56
170 25 -11.87 0.07 -25.49
175 25 -17.86 0.02 -24.38
180 25 -84.56 0.00 -23.25
0 30 -84.95 0.00 -19.57
5 30 -17.86 0.02 -20.48
10 30 -11.87 0.06 -21.40
15 30 -8.40 0.14 -22.31
20 30 -5.97 0.25 -23.20
25 30 -4.13 0.39 -24.07
30 30 -2.66 0.54 -24.92
35 30 -1.45 0.72 -25.73
40 30 -0.45 0.90 -26.51
45 30 0.39 1.09 -27.26
50 30 1.09 1.29 -27.97
55 30 1.69 1.47 -28.65
60 30 2.18 1.65 -29.30
65 30 2.59 1.81 -29.90
70 30 2.91 1.95 -30.47
75 30 3.15 2.07 -30.99
80 30 3.33 2.15 -31.45
85 30 3.43 2.20 -31.87
90 30 3.46 2.22 -32.21
95 30 3.43 2.20 -32.49
100 30 3.33 2.15 -32.69
105 30 3.15 2.07 -32.79
110 30 2.91 1.95 -32.80
115 30 2.59 1.81 -32.71
120 30 2.18 1.65 -32.51
125 30 1.69 1.47 -32.20
130 30 1.09 1.29 -31.79
135 30 0.39 1.09 -31.27
140 30 -0.45 0.90 -30.66
145 30 -1.45 0.72 -29.95
150 30 -2.66 0.54 -29.16
155 30 -4.13 0.39 -28.30
160 30 -5.97 0.25 -27.37
165 30 -8.40 0.14 -26.39
170 30 -11.87 0.07 -25.36
175 30 -17.86 0.02 -24.31
180 30 -84.96 0.00 -23.23
0 35 -85.43 0.00 -19.55
5 35 -17.86 0.02 -20.41
10 35 -11.87 0.06 -21.27
15 35 -8.40 0.14 -22.14
20 35 -5.97 0.25 -22.99
25 35 -4.13 0.39 -23.82
30 35 -2.66 0.54 -24.63
35 35 -1.45 0.72 -25.42
40 35 -0.45 0.90 -26.17
45 35 0.39 1.09 -26.90
50 35 1.09 1.29 -27.58
55 35 1.69 1.47 -28.24
60 35 2.18 1.65 -28.86
65 35 2.59 1.81 -29.44
70 35 2.91 1.95 -29.98
75 35 3.15 2.07 -30.47
80 35 3.33 2.15 -30.91
85 35 3.43 2.20 -31.30
90 35 3.46 2.22 -31.61
95 35 3.43 2.20 -31.87
100 35 3.33 2.15 -32.05
105 35 3.15 2.07 -32.14
110 35 2.91 1.95 -32.15
115 35 2.59 1.81 -32.06
120 35 2.18 1.65 -31.88
125 35 1.69 1.47 -31.60
130 35 1.09 1.29 -31.22
135 35 0.39 1.09 -30.74
140 35 -0.45 0.90 -30.17
145 35 -1.45 0.72 -29.52
150 35 -2.66 0.54 -28.79
155 35 -4.13 0.39 -27.98
160 35 -5.97 0.25 -27.11
165 35 -8.40 0.14 -26.19
170 35 -11.87 0.07 -25.23
175 35 -17.86 0.02 -24.23
180 35 -85.44 0.00 -23.22
0 40 -86.01 0.00 -19.53
... 159721 more lines ...
|
| Attachment 5: XF_model_29_87.txt
|
freq : 83.33 MHz
SWR : 1.965000
Theta Phi Gain(dB) Gain Phase(deg)
0 0 -77.59 0.00 -23.13
5 0 -17.30 0.02 -23.13
10 0 -11.30 0.07 -23.11
15 0 -7.83 0.16 -23.07
20 0 -5.40 0.29 -23.00
25 0 -3.56 0.44 -22.93
30 0 -2.09 0.62 -22.84
35 0 -0.89 0.82 -22.73
40 0 0.11 1.03 -22.61
45 0 0.95 1.24 -22.47
50 0 1.66 1.46 -22.33
55 0 2.25 1.68 -22.17
60 0 2.74 1.88 -22.01
65 0 3.14 2.06 -21.84
70 0 3.46 2.22 -21.66
75 0 3.71 2.35 -21.48
80 0 3.88 2.44 -21.29
85 0 3.99 2.50 -21.11
90 0 4.02 2.52 -20.92
95 0 3.99 2.50 -20.74
100 0 3.88 2.45 -20.55
105 0 3.71 2.35 -20.38
110 0 3.47 2.22 -20.20
115 0 3.15 2.06 -20.03
120 0 2.74 1.88 -19.87
125 0 2.25 1.68 -19.72
130 0 1.66 1.47 -19.58
135 0 0.96 1.25 -19.44
140 0 0.12 1.03 -19.31
145 0 -0.88 0.82 -19.20
150 0 -2.08 0.62 -19.10
155 0 -3.55 0.44 -19.02
160 0 -5.39 0.29 -18.94
165 0 -7.82 0.17 -18.89
170 0 -11.29 0.07 -18.84
175 0 -17.28 0.02 -18.82
180 0 -77.55 0.00 -18.81
0 5 -77.73 0.00 -18.36
5 5 -17.30 0.02 -18.44
10 5 -11.30 0.07 -18.54
15 5 -7.83 0.16 -18.64
20 5 -5.40 0.29 -18.77
25 5 -3.56 0.44 -18.92
30 5 -2.09 0.62 -19.09
35 5 -0.89 0.82 -19.27
40 5 0.11 1.03 -19.47
45 5 0.95 1.24 -19.67
50 5 1.66 1.46 -19.89
55 5 2.25 1.68 -20.12
60 5 2.74 1.88 -20.35
65 5 3.14 2.06 -20.60
70 5 3.46 2.22 -20.84
75 5 3.71 2.35 -21.08
80 5 3.88 2.44 -21.33
85 5 3.99 2.50 -21.57
90 5 4.02 2.52 -21.81
95 5 3.99 2.50 -22.04
100 5 3.88 2.45 -22.28
105 5 3.71 2.35 -22.50
110 5 3.47 2.22 -22.72
115 5 3.15 2.06 -22.92
120 5 2.74 1.88 -23.12
125 5 2.25 1.68 -23.30
130 5 1.66 1.47 -23.47
135 5 0.96 1.25 -23.62
140 5 0.12 1.03 -23.75
145 5 -0.88 0.82 -23.85
150 5 -2.08 0.62 -23.94
155 5 -3.55 0.44 -24.00
160 5 -5.39 0.29 -24.04
165 5 -7.82 0.17 -24.06
170 5 -11.29 0.07 -24.04
175 5 -17.28 0.02 -24.01
180 5 -77.69 0.00 -23.95
0 10 -77.94 0.00 -16.18
5 10 -17.30 0.02 -16.28
10 10 -11.30 0.07 -16.44
15 10 -7.83 0.16 -16.61
20 10 -5.40 0.29 -16.84
25 10 -3.56 0.44 -17.09
30 10 -2.09 0.62 -17.37
35 10 -0.89 0.82 -17.69
40 10 0.11 1.03 -18.03
45 10 0.95 1.24 -18.40
50 10 1.66 1.46 -18.78
55 10 2.25 1.68 -19.19
60 10 2.74 1.88 -19.60
65 10 3.14 2.06 -20.03
70 10 3.46 2.22 -20.46
75 10 3.71 2.35 -20.89
80 10 3.88 2.44 -21.33
85 10 3.99 2.50 -21.76
90 10 4.02 2.52 -22.19
95 10 3.99 2.50 -22.61
100 10 3.88 2.45 -23.02
105 10 3.71 2.35 -23.43
110 10 3.47 2.22 -23.82
115 10 3.15 2.06 -24.20
120 10 2.74 1.88 -24.55
125 10 2.25 1.68 -24.88
130 10 1.66 1.47 -25.19
135 10 0.96 1.25 -25.46
140 10 0.12 1.03 -25.71
145 10 -0.88 0.82 -25.92
150 10 -2.08 0.62 -26.09
155 10 -3.55 0.44 -26.22
160 10 -5.39 0.29 -26.30
165 10 -7.82 0.17 -26.35
170 10 -11.29 0.07 -26.36
175 10 -17.28 0.02 -26.33
180 10 -77.89 0.00 -26.25
0 15 -78.23 0.00 -14.91
5 15 -17.30 0.02 -15.04
10 15 -11.30 0.07 -15.22
15 15 -7.83 0.16 -15.44
20 15 -5.40 0.29 -15.71
25 15 -3.56 0.44 -16.02
30 15 -2.09 0.62 -16.37
35 15 -0.89 0.82 -16.76
40 15 0.11 1.03 -17.19
45 15 0.95 1.24 -17.64
50 15 1.66 1.46 -18.12
55 15 2.25 1.68 -18.63
60 15 2.74 1.88 -19.14
65 15 3.14 2.06 -19.68
70 15 3.46 2.22 -20.21
75 15 3.71 2.35 -20.76
80 15 3.88 2.44 -21.30
85 15 3.99 2.50 -21.85
90 15 4.02 2.52 -22.39
95 15 3.99 2.50 -22.92
100 15 3.88 2.45 -23.44
105 15 3.71 2.35 -23.94
110 15 3.47 2.22 -24.43
115 15 3.15 2.06 -24.91
120 15 2.74 1.88 -25.35
125 15 2.25 1.68 -25.77
130 15 1.66 1.47 -26.16
135 15 0.96 1.25 -26.51
140 15 0.12 1.03 -26.82
145 15 -0.88 0.82 -27.09
150 15 -2.08 0.62 -27.31
155 15 -3.55 0.44 -27.48
160 15 -5.39 0.29 -27.60
165 15 -7.82 0.17 -27.67
170 15 -11.29 0.07 -27.68
175 15 -17.28 0.02 -27.65
180 15 -78.17 0.00 -27.56
0 20 -78.59 0.00 -14.07
5 20 -17.30 0.02 -14.21
10 20 -11.30 0.07 -14.41
15 20 -7.83 0.16 -14.66
20 20 -5.40 0.29 -14.95
25 20 -3.56 0.44 -15.30
30 20 -2.09 0.62 -15.70
35 20 -0.89 0.82 -16.14
40 20 0.11 1.03 -16.61
45 20 0.95 1.24 -17.12
50 20 1.66 1.46 -17.66
55 20 2.25 1.68 -18.23
60 20 2.74 1.88 -18.82
65 20 3.14 2.06 -19.42
70 20 3.46 2.22 -20.03
75 20 3.71 2.35 -20.64
80 20 3.88 2.44 -21.26
85 20 3.99 2.50 -21.88
90 20 4.02 2.52 -22.49
95 20 3.99 2.50 -23.09
100 20 3.88 2.45 -23.69
105 20 3.71 2.35 -24.26
110 20 3.47 2.22 -24.82
115 20 3.15 2.06 -25.36
120 20 2.74 1.88 -25.87
125 20 2.25 1.68 -26.34
130 20 1.66 1.47 -26.78
135 20 0.96 1.25 -27.18
140 20 0.12 1.03 -27.53
145 20 -0.88 0.82 -27.84
150 20 -2.08 0.62 -28.09
155 20 -3.55 0.44 -28.29
160 20 -5.40 0.29 -28.43
165 20 -7.82 0.17 -28.52
170 20 -11.29 0.07 -28.54
175 20 -17.28 0.02 -28.51
180 20 -78.53 0.00 -28.42
0 25 -79.03 0.00 -13.47
5 25 -17.30 0.02 -13.62
10 25 -11.30 0.07 -13.83
15 25 -7.83 0.16 -14.09
20 25 -5.40 0.29 -14.41
25 25 -3.56 0.44 -14.78
30 25 -2.09 0.62 -15.20
35 25 -0.89 0.82 -15.67
40 25 0.11 1.03 -16.18
45 25 0.95 1.24 -16.73
50 25 1.66 1.46 -17.31
55 25 2.25 1.68 -17.92
60 25 2.74 1.88 -18.55
65 25 3.14 2.06 -19.20
70 25 3.46 2.22 -19.86
75 25 3.71 2.35 -20.53
80 25 3.88 2.44 -21.20
85 25 3.99 2.50 -21.87
90 25 4.02 2.52 -22.54
95 25 3.99 2.50 -23.20
100 25 3.88 2.45 -23.84
105 25 3.71 2.35 -24.47
110 25 3.47 2.22 -25.08
115 25 3.15 2.06 -25.66
120 25 2.74 1.88 -26.21
125 25 2.25 1.68 -26.73
130 25 1.66 1.47 -27.21
135 25 0.96 1.25 -27.64
140 25 0.12 1.03 -28.03
145 25 -0.88 0.82 -28.36
150 25 -2.08 0.62 -28.64
155 25 -3.55 0.44 -28.86
160 25 -5.40 0.29 -29.02
165 25 -7.82 0.17 -29.12
170 25 -11.29 0.07 -29.15
175 25 -17.28 0.02 -29.13
180 25 -78.97 0.00 -29.04
0 30 -79.57 0.00 -13.01
5 30 -17.30 0.02 -13.16
10 30 -11.30 0.07 -13.37
15 30 -7.83 0.16 -13.65
20 30 -5.40 0.29 -13.97
25 30 -3.56 0.44 -14.36
30 30 -2.09 0.62 -14.80
35 30 -0.89 0.82 -15.29
40 30 0.11 1.03 -15.82
45 30 0.95 1.24 -16.40
50 30 1.66 1.46 -17.01
55 30 2.25 1.68 -17.65
60 30 2.74 1.88 -18.32
65 30 3.14 2.06 -19.00
70 30 3.46 2.22 -19.70
75 30 3.71 2.35 -20.41
80 30 3.88 2.44 -21.12
85 30 3.99 2.50 -21.84
90 30 4.02 2.52 -22.55
95 30 3.99 2.50 -23.24
100 30 3.88 2.45 -23.93
105 30 3.71 2.35 -24.60
110 30 3.47 2.22 -25.25
115 30 3.15 2.06 -25.87
120 30 2.74 1.88 -26.46
125 30 2.25 1.68 -27.01
130 30 1.66 1.47 -27.52
135 30 0.96 1.25 -27.98
140 30 0.12 1.03 -28.39
145 30 -0.88 0.82 -28.75
150 30 -2.08 0.62 -29.05
155 30 -3.55 0.44 -29.28
160 30 -5.40 0.29 -29.46
165 30 -7.82 0.17 -29.57
170 30 -11.29 0.07 -29.61
175 30 -17.28 0.02 -29.59
180 30 -79.50 0.00 -29.50
0 35 -80.22 0.00 -12.65
5 35 -17.30 0.02 -12.80
10 35 -11.30 0.07 -13.01
15 35 -7.83 0.16 -13.28
20 35 -5.40 0.29 -13.62
25 35 -3.56 0.44 -14.02
30 35 -2.09 0.62 -14.47
35 35 -0.89 0.82 -14.97
40 35 0.11 1.03 -15.52
45 35 0.95 1.24 -16.11
50 35 1.66 1.46 -16.75
55 35 2.25 1.68 -17.41
60 35 2.74 1.88 -18.10
65 35 3.14 2.06 -18.81
70 35 3.46 2.22 -19.54
75 35 3.71 2.35 -20.28
80 35 3.88 2.44 -21.03
85 35 3.99 2.50 -21.78
90 35 4.02 2.52 -22.52
95 35 3.99 2.50 -23.26
100 35 3.88 2.45 -23.98
105 35 3.71 2.35 -24.68
110 35 3.47 2.22 -25.36
115 35 3.15 2.06 -26.02
120 35 2.74 1.88 -26.63
125 35 2.25 1.68 -27.21
130 35 1.66 1.47 -27.75
135 35 0.96 1.25 -28.23
140 35 0.12 1.03 -28.66
145 35 -0.88 0.82 -29.04
150 35 -2.08 0.62 -29.36
155 35 -3.55 0.44 -29.61
160 35 -5.40 0.29 -29.80
165 35 -7.82 0.17 -29.92
170 35 -11.29 0.07 -29.97
175 35 -17.28 0.02 -29.96
180 35 -80.14 0.00 -29.87
0 40 -80.99 0.00 -12.34
... 159721 more lines ...
|
|
2
|
Wed Feb 6 17:28:29 2019 |
Julie Rolla | Bicone User Manual (w/ links to Dipole work) | Hi all,
The working copy of the user manual can be found here: https://www.overleaf.com/6955763886pjtkckprxsyp
It still need to be tested by someone new to see if edits need to be made. I have uploaded the copy that is finalized as of this day. Note if you click on the overleaf version that may be more up-to-date. This copy is added as a PDF so we can keep track of versions. |
| Attachment 1: GENETIS_Manual_1_25_19.pdf
|
|
17
|
Thu Nov 7 16:55:00 2019 |
Cade Sbrocco | Archiving Old Genetis Manual | |
| Attachment 1: GENETIS_Manual.pdf
|
|
11
|
Wed Apr 3 16:38:07 2019 |
Julie Rolla | AraSim update | General update
The following were the fixes we made to make AraSim run:
(1) As of March, the loop breaks at AraSim. We have found that our program XFintoARA.py does not properly format the output in the form that AraSim is looking for (as an input). Jorge gave us a sample AraSim input file to help us edit XFintoARA.py so that it formats the file correctly. Note that we found white space is also must match in order for the format to be correct.
Now that we have fixed this, AraSim now runs. Attached is the following sample Jorge gave us of the proper AraSim inputs. If needed, it's available to refer back to.
(2) After correcting the AraSim input file formats (outputs of XFintoARA.py), we ran it. It failed to finish running after hours of run time. We found that we were using default input parameters for AraSim -- which wasn't triggering in a timely fashion. Once we edited the parameters, it ran successfully. Note that this was not run within the loop. Confirmation of the full loop running has not been completed.
The parameters of interest need to be edited in setup.txt. To see which file in AraSim I am referring to, please see the .txt file attached. Note that we will be editing the bash script to allow the user to change these input parameters there. The following red lines are what need to be set:
EXPONENT=21
NNU=100 // number of neutrino events
NNU_PASSED=10 // number of neutrino events that are allowed to pass the trigger
ONLY_PASSED_EVENTS=1 // 0 (default): AraSim throws NNU events whether or not they pass; 1: AraSim throws events until the number of events that pass the trigger is equal to NNU_PASSED (WARNING: may cause long run times if reasonable values are not chosen)
NOISE_WAVEFORM_GENERATE_MODE=0 // generate new noise waveforms for each events
NOISE_EVENTS=16 // number of pure noise waveforms
TRIG_ANALYSIS_MODE=0 // 0 = signal + noise, 1 = signal only, 2 = noise only
DETECTOR=1 // ARA stations 1 to 7
NOFZ=1
core_x=10000
core_y=10000
EXPONENT gives the energy of the neutrinos. Note that higher energies will have more pass, as the energies don't dissipate as rapidly. For now, we are running at this energy. Eventally, we will want to use the full Kotera spectrum. Otherwsie, we would be finding the performance of an antenna at only one energy. Just to check the loop, we are going to use EXPONENT=21 to get events to trigger faster.
ONLY_PASSED_EVENTS decides which of the two options to use to determine when AraSim is finished running: (1) Throws 'NNU' number of events at antennas and stops regardless of how many pass (in this case ONLY_PASSED_EVENTS=0) , or (2) throws as many events at the antenna as needed until 'NNU_PASSED' number triggers (in this case ONLY_PASSED_EVENTS=1). Ie if this number is 10, we throw as many neutrinos at the antenna as necessary until 10 events trigger. Note that this can result in a longer run time. Additionally, a higher EXPONENT will make them trigger faster.
TRIG_ANALYSIS_MODE is a setting that determines what is being thrown. Ie- 0=signal + noise (which is what we want, and is the 'real' case), 1=signal only (would make it run faster), 2=noise only (not of interest for us).
To-Do List
(1) Commit version of XFintoARA.py since recent formatting change.
(2) Figure out how to add in Kotera spectrum (note that we wont want to do this until the WHOLE loop is functional)
(3) Edit the bash script to recreat the setup.txt making the variables in red above an editable option in the top of the bash script. Ie our variables in red above will be at the top of the bash script. Later in the bash script we will need to have it create this .txt file with the user inputs for those variables. We can do this by either (1) creating a skeleton, or (2) creating another .sh script that is run in our XFLoop.sh -- which writes the first handfull of lines based on the user inputs for our variables in red above, and then pastes in the rest of what is unchanged in setup.txt.
(4) Run the whole loop: update- we tried this today and keep getting a Veff-0 for our first antenna along with the error
Warning in <TFile::Init>: no StreamerInfo found in /datapool/software/anita/lib/G__RootFftwWrapper_rdict.pcm therefore preventing schema evolution when reading this file.
Error in <TStreamerInfo::Build>: AraEventCalibrator, discarding: unsigned short* fAtriPeds, no [dimension]
others run as expected
(5) Work on the paper: Amy added a skeleton of sections to add for the GA. Date of completion should be in-line with APS.
(6) APS talk slides.
(7) Edit the manual and add:
- The changes that will occur when task (3) is done.
- A sample of how AraSim inputs -- or XFintoARA.py outputs -- should look.
- This stuff: http://radiorm.physics.ohio-state.edu/elog/GENETIS/8
- Max's AraSim plotting software (when it's done)
- Evelyn and Sophie's plotting software info (when it's done).
(8) Finish Max's AraSim plotting software.
(9) Finish Evenlyn and Sophie's plotting softare. |
| Attachment 1: ARA_bicone6in_output.txt
|
freq : 83.33 MHz
SWR : 2.18
Theta Phi Gain(dB) Gain Phase(deg)
0 0 -80.83 0.00 -0.41
5 0 -21.10 0.01 168.18
10 0 -15.10 0.03 168.17
15 0 -11.62 0.07 168.14
20 0 -9.19 0.12 168.09
25 0 -7.34 0.18 168.04
30 0 -5.86 0.26 167.97
35 0 -4.65 0.34 167.91
40 0 -3.64 0.43 167.83
45 0 -2.79 0.53 167.76
50 0 -2.08 0.62 167.70
55 0 -1.48 0.71 167.64
60 0 -0.98 0.80 167.59
65 0 -0.57 0.88 167.55
70 0 -0.25 0.94 167.53
75 0 0.00 1.00 167.51
80 0 0.18 1.04 167.50
85 0 0.29 1.07 167.50
90 0 0.32 1.08 167.50
95 0 0.29 1.07 167.50
100 0 0.18 1.04 167.50
105 0 0.01 1.00 167.51
110 0 -0.25 0.94 167.53
115 0 -0.57 0.88 167.55
120 0 -0.98 0.80 167.59
125 0 -1.48 0.71 167.64
130 0 -2.08 0.62 167.70
135 0 -2.79 0.53 167.77
140 0 -3.64 0.43 167.84
145 0 -4.65 0.34 167.91
150 0 -5.86 0.26 167.98
155 0 -7.33 0.18 168.04
160 0 -9.19 0.12 168.10
165 0 -11.62 0.07 168.15
170 0 -15.09 0.03 168.18
175 0 -21.08 0.01 168.21
180 0 -80.83 0.00 179.59
0 5 -80.83 0.00 -0.41
5 5 -21.10 0.01 168.18
10 5 -15.10 0.03 168.17
15 5 -11.62 0.07 168.14
20 5 -9.19 0.12 168.09
25 5 -7.34 0.18 168.04
30 5 -5.86 0.26 167.97
35 5 -4.65 0.34 167.91
40 5 -3.64 0.43 167.83
45 5 -2.79 0.53 167.76
50 5 -2.08 0.62 167.70
55 5 -1.48 0.71 167.64
60 5 -0.98 0.80 167.59
65 5 -0.57 0.88 167.55
70 5 -0.25 0.94 167.53
75 5 0.00 1.00 167.51
80 5 0.18 1.04 167.50
85 5 0.29 1.07 167.50
90 5 0.32 1.08 167.50
95 5 0.29 1.07 167.50
100 5 0.18 1.04 167.50
105 5 0.01 1.00 167.51
110 5 -0.25 0.94 167.53
115 5 -0.57 0.88 167.56
120 5 -0.98 0.80 167.59
125 5 -1.48 0.71 167.64
130 5 -2.08 0.62 167.70
135 5 -2.79 0.53 167.77
140 5 -3.64 0.43 167.84
145 5 -4.65 0.34 167.91
150 5 -5.86 0.26 167.98
155 5 -7.33 0.18 168.04
160 5 -9.19 0.12 168.10
165 5 -11.62 0.07 168.15
170 5 -15.09 0.03 168.18
175 5 -21.08 0.01 168.21
180 5 -80.83 0.00 179.59
0 10 -80.83 0.00 -0.41
5 10 -21.10 0.01 168.18
10 10 -15.10 0.03 168.17
15 10 -11.62 0.07 168.14
20 10 -9.19 0.12 168.09
25 10 -7.34 0.18 168.04
30 10 -5.86 0.26 167.97
35 10 -4.65 0.34 167.91
40 10 -3.64 0.43 167.83
45 10 -2.79 0.53 167.76
50 10 -2.08 0.62 167.70
55 10 -1.48 0.71 167.64
60 10 -0.98 0.80 167.59
65 10 -0.57 0.88 167.55
70 10 -0.25 0.94 167.53
75 10 0.01 1.00 167.51
80 10 0.18 1.04 167.50
85 10 0.29 1.07 167.50
90 10 0.32 1.08 167.50
95 10 0.29 1.07 167.50
100 10 0.18 1.04 167.50
105 10 0.01 1.00 167.51
110 10 -0.24 0.95 167.53
115 10 -0.57 0.88 167.56
120 10 -0.98 0.80 167.59
125 10 -1.48 0.71 167.64
130 10 -2.08 0.62 167.70
135 10 -2.79 0.53 167.77
140 10 -3.64 0.43 167.84
145 10 -4.65 0.34 167.91
150 10 -5.86 0.26 167.98
155 10 -7.33 0.18 168.04
160 10 -9.19 0.12 168.10
165 10 -11.62 0.07 168.15
170 10 -15.09 0.03 168.18
175 10 -21.08 0.01 168.21
180 10 -80.83 0.00 179.59
0 15 -80.83 0.00 -0.41
5 15 -21.10 0.01 168.18
10 15 -15.10 0.03 168.17
15 15 -11.62 0.07 168.14
20 15 -9.19 0.12 168.09
25 15 -7.34 0.18 168.04
30 15 -5.86 0.26 167.97
35 15 -4.65 0.34 167.91
40 15 -3.64 0.43 167.83
45 15 -2.79 0.53 167.76
50 15 -2.08 0.62 167.70
55 15 -1.48 0.71 167.64
60 15 -0.98 0.80 167.59
65 15 -0.57 0.88 167.55
70 15 -0.25 0.94 167.53
75 15 0.01 1.00 167.51
80 15 0.18 1.04 167.50
85 15 0.29 1.07 167.50
90 15 0.32 1.08 167.50
95 15 0.29 1.07 167.50
100 15 0.18 1.04 167.50
105 15 0.01 1.00 167.51
110 15 -0.24 0.95 167.53
115 15 -0.57 0.88 167.56
120 15 -0.98 0.80 167.59
125 15 -1.48 0.71 167.64
130 15 -2.08 0.62 167.70
135 15 -2.79 0.53 167.77
140 15 -3.64 0.43 167.84
145 15 -4.65 0.34 167.91
150 15 -5.86 0.26 167.98
155 15 -7.33 0.18 168.04
160 15 -9.19 0.12 168.10
165 15 -11.62 0.07 168.15
170 15 -15.09 0.03 168.18
175 15 -21.08 0.01 168.21
180 15 -80.83 0.00 179.59
0 20 -80.83 0.00 -0.41
5 20 -21.10 0.01 168.18
10 20 -15.10 0.03 168.17
15 20 -11.62 0.07 168.14
20 20 -9.19 0.12 168.09
25 20 -7.34 0.18 168.04
30 20 -5.86 0.26 167.97
35 20 -4.65 0.34 167.91
40 20 -3.64 0.43 167.83
45 20 -2.79 0.53 167.76
50 20 -2.08 0.62 167.70
55 20 -1.48 0.71 167.64
60 20 -0.98 0.80 167.59
65 20 -0.57 0.88 167.55
70 20 -0.24 0.95 167.53
75 20 0.01 1.00 167.51
80 20 0.18 1.04 167.50
85 20 0.29 1.07 167.50
90 20 0.32 1.08 167.50
95 20 0.29 1.07 167.50
100 20 0.18 1.04 167.50
105 20 0.01 1.00 167.51
110 20 -0.24 0.95 167.53
115 20 -0.57 0.88 167.56
120 20 -0.98 0.80 167.59
125 20 -1.48 0.71 167.64
130 20 -2.08 0.62 167.70
135 20 -2.79 0.53 167.77
140 20 -3.64 0.43 167.84
145 20 -4.65 0.34 167.91
150 20 -5.86 0.26 167.98
155 20 -7.33 0.18 168.04
160 20 -9.19 0.12 168.10
165 20 -11.62 0.07 168.15
170 20 -15.09 0.03 168.18
175 20 -21.08 0.01 168.21
180 20 -80.83 0.00 179.59
0 25 -80.83 0.00 -0.41
5 25 -21.10 0.01 168.18
10 25 -15.10 0.03 168.17
15 25 -11.62 0.07 168.14
20 25 -9.19 0.12 168.09
25 25 -7.34 0.18 168.04
30 25 -5.86 0.26 167.97
35 25 -4.65 0.34 167.91
40 25 -3.64 0.43 167.83
45 25 -2.79 0.53 167.76
50 25 -2.08 0.62 167.70
55 25 -1.48 0.71 167.64
60 25 -0.98 0.80 167.59
65 25 -0.57 0.88 167.55
70 25 -0.24 0.95 167.53
75 25 0.01 1.00 167.51
80 25 0.18 1.04 167.50
85 25 0.29 1.07 167.50
90 25 0.32 1.08 167.50
95 25 0.29 1.07 167.50
100 25 0.18 1.04 167.50
105 25 0.01 1.00 167.51
110 25 -0.24 0.95 167.53
115 25 -0.57 0.88 167.56
120 25 -0.98 0.80 167.59
125 25 -1.48 0.71 167.64
130 25 -2.08 0.62 167.70
135 25 -2.79 0.53 167.77
140 25 -3.64 0.43 167.84
145 25 -4.65 0.34 167.91
150 25 -5.86 0.26 167.98
155 25 -7.33 0.18 168.04
160 25 -9.19 0.12 168.10
165 25 -11.62 0.07 168.15
170 25 -15.09 0.03 168.18
175 25 -21.08 0.01 168.21
180 25 -80.83 0.00 179.59
0 30 -80.83 0.00 -0.41
5 30 -21.10 0.01 168.19
10 30 -15.10 0.03 168.17
15 30 -11.62 0.07 168.14
20 30 -9.19 0.12 168.09
25 30 -7.34 0.18 168.04
30 30 -5.86 0.26 167.98
35 30 -4.65 0.34 167.91
40 30 -3.64 0.43 167.83
45 30 -2.79 0.53 167.76
50 30 -2.08 0.62 167.70
55 30 -1.48 0.71 167.64
60 30 -0.98 0.80 167.59
65 30 -0.57 0.88 167.55
70 30 -0.24 0.95 167.53
75 30 0.01 1.00 167.51
80 30 0.18 1.04 167.50
85 30 0.29 1.07 167.50
90 30 0.32 1.08 167.50
95 30 0.29 1.07 167.50
100 30 0.18 1.04 167.51
105 30 0.01 1.00 167.51
110 30 -0.24 0.95 167.53
115 30 -0.57 0.88 167.56
120 30 -0.98 0.80 167.59
125 30 -1.48 0.71 167.64
130 30 -2.08 0.62 167.70
135 30 -2.79 0.53 167.77
140 30 -3.64 0.43 167.84
145 30 -4.65 0.34 167.91
150 30 -5.86 0.26 167.98
155 30 -7.33 0.18 168.04
160 30 -9.19 0.12 168.10
165 30 -11.62 0.07 168.15
170 30 -15.09 0.03 168.18
175 30 -21.08 0.01 168.21
180 30 -80.83 0.00 179.59
0 35 -80.83 0.00 -0.41
5 35 -21.10 0.01 168.19
10 35 -15.10 0.03 168.17
15 35 -11.62 0.07 168.14
20 35 -9.19 0.12 168.09
25 35 -7.34 0.18 168.04
30 35 -5.86 0.26 167.98
35 35 -4.65 0.34 167.91
40 35 -3.64 0.43 167.83
45 35 -2.79 0.53 167.76
50 35 -2.08 0.62 167.70
55 35 -1.48 0.71 167.64
60 35 -0.98 0.80 167.59
65 35 -0.57 0.88 167.56
70 35 -0.24 0.95 167.53
75 35 0.01 1.00 167.51
80 35 0.18 1.04 167.50
85 35 0.29 1.07 167.50
90 35 0.32 1.08 167.50
95 35 0.29 1.07 167.50
100 35 0.18 1.04 167.51
105 35 0.01 1.00 167.51
110 35 -0.24 0.95 167.53
115 35 -0.57 0.88 167.56
120 35 -0.98 0.80 167.59
125 35 -1.48 0.71 167.64
130 35 -2.08 0.62 167.70
135 35 -2.79 0.53 167.77
140 35 -3.64 0.43 167.84
145 35 -4.65 0.34 167.91
150 35 -5.86 0.26 167.98
155 35 -7.33 0.18 168.04
160 35 -9.18 0.12 168.10
165 35 -11.62 0.07 168.15
170 35 -15.09 0.03 168.18
175 35 -21.08 0.01 168.21
180 35 -80.83 0.00 179.59
0 40 -80.83 0.00 -0.41
... 159721 more lines ...
|
| Attachment 2: setup.txt
|
///////////////////
NFOUR=1024
EXPONENT=21
NNU=100 // number of neutrino events
NNU_PASSED=10 // number of neutrino events that are allowed to pass the trigger
ONLY_PASSED_EVENTS=1 // 0 (default): AraSim throws NNU events whether or not they pass; 1: AraSim throws events until the number of events that pass the trigger is equal to NNU_PASSED (WARNING: may cause long run times if reasonable values are not chosen)
NOISE_WAVEFORM_GENERATE_MODE=0 // generate new noise waveforms for each events
NOISE_EVENTS=16 // number of pure noise waveforms
TRIG_ANALYSIS_MODE=0 // 0 = signal + noise, 1 = signal only, 2 = noise only
DETECTOR=1 // ARA stations 1 to 7
NOFZ=1
core_x=10000
core_y=10000
TIMESTEP=5.E-10 // value for 2GHz actual station value
TRIG_WINDOW=1.E-7 // 100ns which is actual testbed trig window
POWERTHRESHOLD=-6.06 // 100Hz global trig rate for 3 out of 16 ARA stations
//DATA_BIN_SIZE=8192
POSNU_RADIUS=3000
V_MIMIC_MODE=0 // 0 : global trig is located center of readout windows
DATA_SAVE_MODE=0 // 2 : don't save any waveform informations at all
DATA_LIKE_OUTPUT=0 // 0 : don't save any waveform information to eventTree
BORE_HOLE_ANTENNA_LAYOUT=0
SECONDARIES=0
// below settings are available for only TestBed mode (DETECTOR=3 case)
TRIG_ONLY_BH_ON=0
CALPULSER_ON=0
USE_MANUAL_GAINOFFSET=0
USE_TESTBED_RFCM_ON=0
NOISE_TEMP_MODE=0
TRIG_THRES_MODE=0
READGEOM=0 // reads geometry information from the sqlite file or not (0 : don't read)
// new variables
//INTERACTION_MODE=0 // pickunbiased mode!
//taumodes=1
//BH_ANT_SEP_DIST_ON=0 // default 0 : use constant borehole antenna distance. 1 : use separate antenna distance. use z_btw01, z_btw12, ... in ARA_N_info.txt or ARA37_info.txt
TRIG_MODE=0 // use vpol, hpol separated trigger mode. by default N_TRIG_V=3, N_TRIG_H=3. You can change this values
//N_TRIG_V=4
//N_TRIG_H=4
number_of_stations=1
core_x=10000
core_y=10000
///////////////////
//DATA_SAVE_MODE=2
// Settings for Arbitrary event generation
//EVENT_TYPE=10 // arbitrary event type
//INTERACTION_MODE=2 // pick exact location using settings below
//POSNU_THETA=1.7 // interaction location elevation angle in radians
//POSNU_THETA=-0.5 // interaction location elevation angle in radians
//POSNU_PHI=0 // interaction location azimuthal angle in radians
//POSNU_R=1000 // interaction location radius in meters
//INTERACTION_MODE=4 // pick interaction location within cylindrical volume above the ice
//PICK_ABOVE_HEIGHT=3000.
// Settings for Station A2-like simulation
//DETECTOR=4
//DETECTOR_STATION=2
//DATA_LIKE_OUTPUT=1
///////////////////////// Other Alternate Settings //////////////////
//POSNU_RADIUS=5000
//RAYSOL_RANGE=7000
//SELECT_FLAVOR=1
//NU_NUBAR_SELECT_MODE=1
//SELECT_CURRENT=1 //default 2: get from ratios in Ghandi etal paper, 0: nc, 1: cc
//NNU_THIS_THETA=1
//NNU_THETA=0.785
//NNU_D_THETA=0.0873
//NNU_D_THETA=0.0
//NOISE_EVENTS=16
//NOISE_EVENTS=1
//NOISE=0
//NOISE_TEMP_MODE=1
//ANTENNA_MODE=1
//APPLY_NOISE_FIGURE=1
//ALL_ANT_V_ON=0
|
|
197
|
Mon Feb 20 12:33:20 2023 |
Dennis H. Calderon | AraSim Simulations for Top 5 Antennas (12/2022) with Higher Stats | AraSim Simulations for Top 5 Antennas from 12/2022 Run with 3 million simulated events
Simulated using Arasim version currently used in GENETIS Loop
/fs/ess/PAS1960/BiconeEvolutionOSC/AraSim/
AraSim ROOT files stored in
/fs/ess/PAS1960/ROOT_Files_Higher_Stats/AraSim_Loop_122022/Feb_2023_Crazy_Sides/
Antennas simulated
- Generation: 13, Individual: 84
- Generation: 18, Individual: 89
- Generation: 19, Individual: 96
- Generation: 29, Individual: 87
- Generation: 9, Individual: 50
Results
| Antenna |
Total Events |
Triggered Events |
Weighted Events |
Effective Volume |
Effective Volume Error |
| 13_84 |
2995982 |
40251 |
16058 |
5.71 |
0.035 |
| 18_89 |
3010000 |
40224 |
15963 |
5.65 |
0.035 |
| 19_96 |
3010000 |
39989 |
16034 |
5.68 |
0.031 |
| 29_87 |
3010000 |
40056 |
16049 |
5.68 |
0.034 |
| 9_50 |
3010000 |
40042 |
16057 |
5.69 |
0.034 |
| ARA_Sym_Bicone_6in |
3010000 |
36627 |
14334 |
5.08 |
0.032 |
Conclusion
Antenna from Generation 13 Individual 84 performed the best with these stats |
| Attachment 1: Crazy_Sides_Triggered_Event_Data.pdf
|
|
240
|
Tue May 21 09:55:57 2024 |
Jacob Weiler | AraSim CSE Spring 2024 Work | # AraSim CSE Spring 2024 Work
## Goals
The main goal was to get a working multithreaded version of the AraSim codebase working. Doing this, the hope was to learn how to multithread the code and get it in a good place to hopefully also integrate GPU's at a later date.
## Where is it at currently?
A bulk of the work done was to functionalize the Connect_Interaction_Detector_V2 to allow for multithreading and to cleanup codebase.
### Going through code added/changed
Helper functions for multiple parts of code. They went through and split it up into multiple parts. Putting line numbers when needed
Part 1: Clearning Antenna Data
Part 2: Determine gain channel
Part 3: solve ray tracing (Not Done)
Part 4: Process Ray Tracing Solution (Not Done)
Part 5: Calculate Signal Factors (Not Done)
Part 6: Calculate Antenna Gain Factors (Not Done)
Part 7: Process Frequency Domain Signal
Part 8A: Process Neutrino Events. Lines 908 - 1208 (Not Done)
Part 8B: Process Arbitrary Events. Lines 1209 - 1475
Part 8C: Process Simple Pulser Simulation. Lines 1478 - 1747 (Not Done)
Part 8D: Process PVA Pulser Simulation. Lines 1751 - 2110 (Not Done)
Part 8E: Process Calpulser Event. Lines 2113 - 2445
Part 9A: Process Noise. Lines 2593 - 2955
Part 9B: Process Trigger and Mimic Waveforms. Lines 2994 - 3624
## What still needs to be done?
- Multithreading still isn't working, multiple threads are writing data to the same place causing the program to crash. This need to be resolved to at least have a working prototype.
- I believe for multithreading we need to mark explicitly where file/data writing is happening to be able to adjust to make thread-safe
- Double checking that new functions are passing variables in the correct way. The CSE students had this has a slight fear.
- Some helper functions are still not completed (3-6, 8A, 8C, 8D)
- Current completed parts of code are all in separate branches and need to be merged after double checking that variable passing is correct |
|
148
|
Fri Mar 25 17:14:01 2022 |
Alex M | Antenna Minimum Length Investigation | The reviewer of the paper we are trying to publish (as well as other external colleagues we showed the paper to) asked about our minimum length we implemented in the loop. Currently, we cut off the length at a minimum of 37.5 cm for each cone (minimum of 75 cm for the full cone). There is an ELOG post from Amy that presented the reasoning for this ( http://radiorm.physics.ohio-state.edu/elog/GENETIS/81 ). We initially thought that we needed a minimum because the antennas were evolving to be small (around and lower than the minimum). We thought that XFdtd was being inaccurate at lengths lower than 37.5 cm, so we set that as a minimum. To see if we can replicate any strange behavior from XFdtd, I simulated antennas at and below our minimum length and generated plots of their antennas responses. The genes are listed below, corresponding to the patterns in the plot. The results do not look unusual (that is, not dissimilar to the other bicones we have generated, including the ARA bicone) and seem to show an improved sensitivity in the upward direction for shorter bicones. The equation we used to arrive at this minimum (f = c/(4L)) might be indicative of a maximum length rather than a minimum.
| Inner radius |
Length |
Quadratic |
Linear |
2.4892,37.5,-0.00142604,0.032832
4.64522,37.5,0.00153863,-0.14004
2.4892,34.5,-0.00142604,0.032832
4.64522,34.5,0.00153863,-0.14004
2.4892,31.5,-0.00142604,0.032832
4.64522,31.5,0.00153863,-0.14004
2.4892,27.5,-0.00142604,0.032832
4.64522,27.5,0.00153863,-0.14004
2.4892,23.5,-0.00142604,0.032832
4.64522,23.5,0.00153863,-0.14004
2.4892,19.5,-0.00142604,0.032832
4.64522,19.5,0.00153863,-0.14004
2.4892,15.5,-0.00142604,0.032832
4.64522,15.5,0.00153863,-0.14004
|
| Attachment 1: polar_plot_283.37.png
| 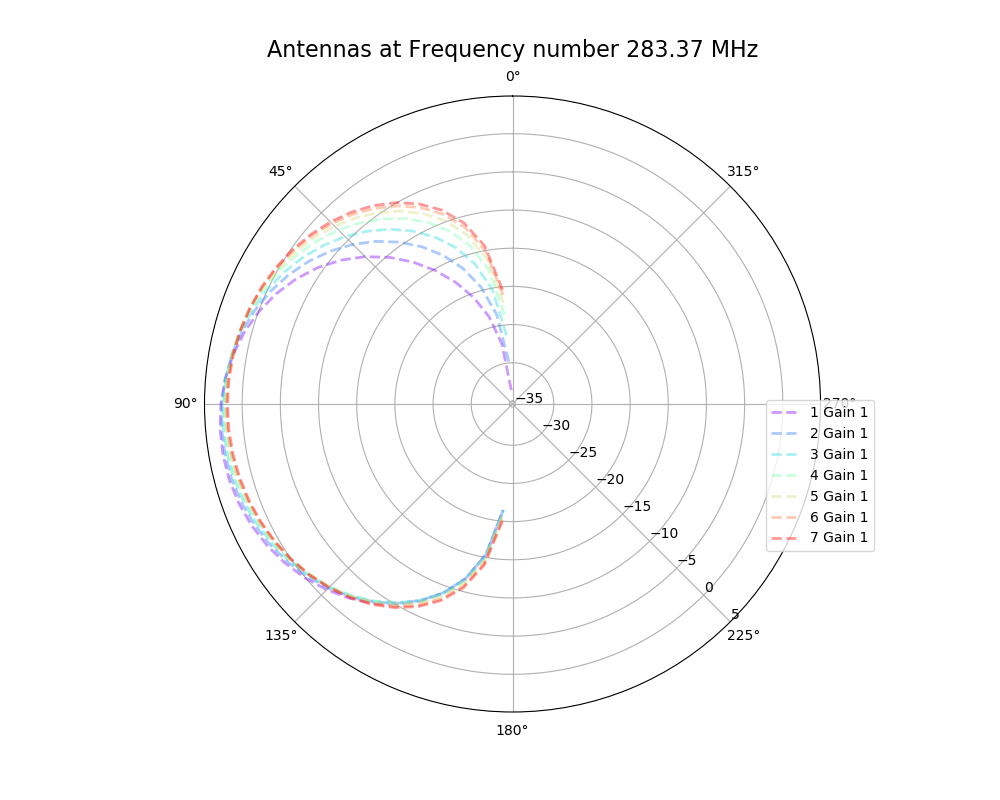 |
|
27
|
Thu Dec 5 14:32:54 2019 |
Julie Rolla | Adding Constraints To-Do List | In my last post, I mentioned (1) our to-do list for adding the ARA hole constraints, and (2) that we've made a mistake upon implementing it. I've attached detailed steps of that to-do list that I've written, as well as an explanation for the correction that is currently being worked on. Please see the pdf attached for an explanation of corrections.
|
| Attachment 1: AddingConstraints.pdf
|
|
Draft
|
Tue Apr 11 00:36:49 2023 |
Bryan Reynolds | AREA physics of results plot first look | After some work, I've gotten the physics of results plotting script provided by Dennis working to compare the ARASim outputs from an AREA run to a set of reference data for the ARA bicone (provided by Alex). Among the outputs is a histogram of of events triggered vs cos(theta_nu) (attached here). This is the plot that was previously presented in GENETIS talks in the physics of results discussion.
Here I compared the best individual from the AREA results (Blue- "source_2") that I have on hand (only the ARASim output files from the most recent generation ran are currently saved) with the ARA bicone reference data (Red- "source_1"). Both datasets included a total of 150,000 NNU. Note that the AREA run used was a single frequency test, meaning that the radiation pattern was held constant for all frequencies in the band.
Also attached for reference are the gain pattern of the associated individual from the AREA output and the violin plot of the run producing the data. |
| Attachment 1: 20230410_AREA_PoR_CosTheta_bestIndiv_singleFreqTest.PNG
| 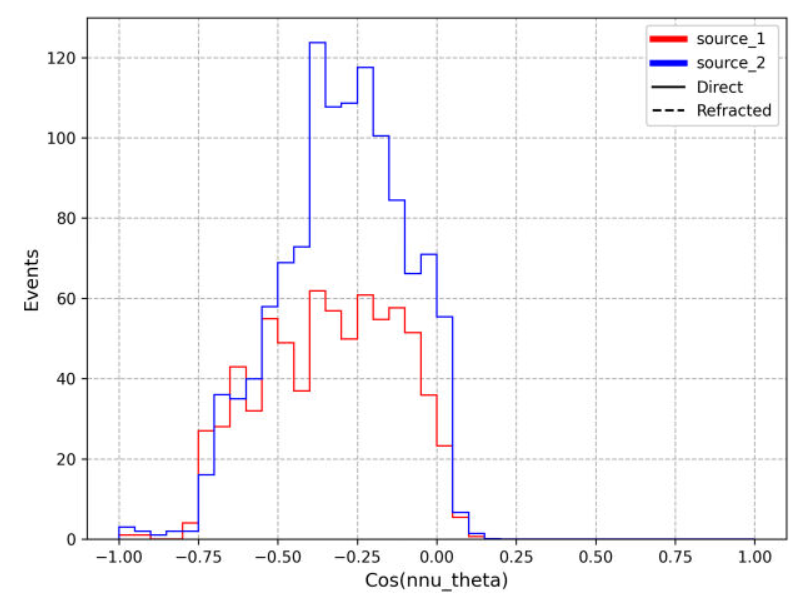 |
| Attachment 2: 20230201_singleFreqTest_100NPop_fitnessPlot_10Gens.PNG
| 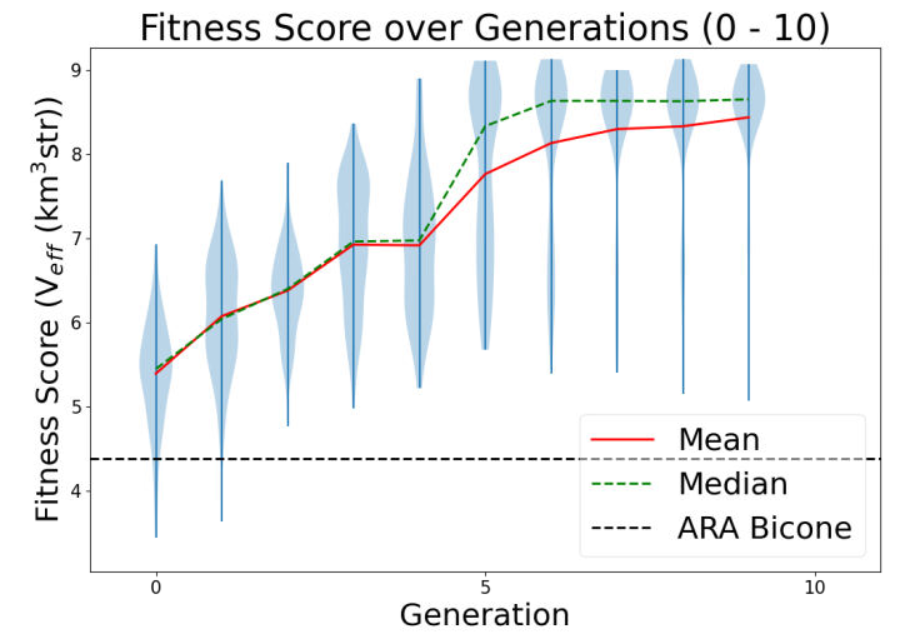 |
| Attachment 3: 20230410_AREA_mostFitIndivReUpload_singleFreqTest.PNG
| 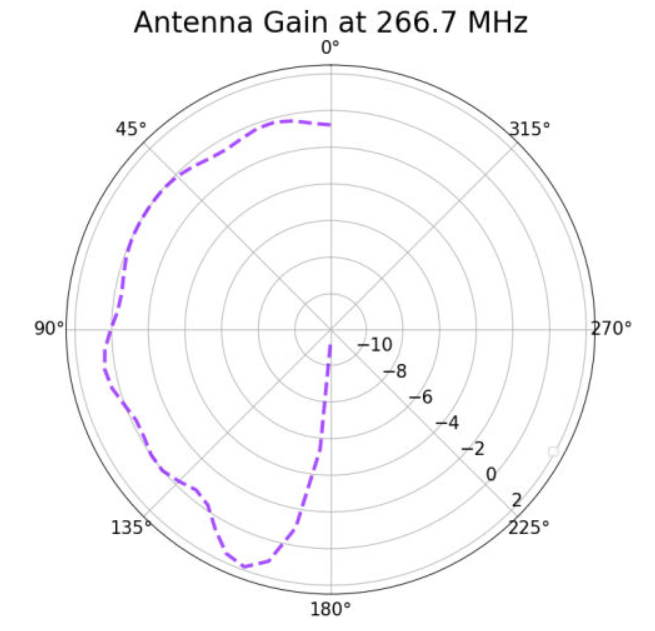 |
|
218
|
Thu May 11 15:57:08 2023 |
Ryan Debolt, Byran Reynolds | AREA Updates | Here is some backlogged information as well as recent updates to our progress on AREA and its optimizations:
4/13/2023
We concluded our initial test of the AREA optimization loop. While analyzing our results, we noticed that most of our runs never reached our benchmarks for performance (chi-squared scores of .25 and .1) and that those that did contain abnormally high amounts of mutation. From previous runs using the GA for the GENETIS loop, we find that mutation and immigration (Note: AREA does not use immigration) usually play a smaller role in overall growth and are simply there to promote diversity. Crossover on the other hand is what handles the bulk of the growth done by the GA. Given our results ran counter to that, we decided to look further into the matter.
Upon looking at the fitness score files, we could see that mutation-created individuals did not perform well in the best run types, but the crossover ones did contain the bulk of good scores. This lines up with expectations. However, closer inspection revealed that these crossover-derived individuals had extremely similar fitness scores, which could indicate elitism in selection methods leading to a lack of diversity in solutions. We then looked at a run that used only crossover with one selection type (roulette), and found every individual to have identical (and poor) scores that were the same up to around the 4th decimal place. This is a strong case that the selection methods used are too elitist to grow a population properly.
We dove into the AREA GA to take a look at the functions doing the selection methods, and discovered several issues potentially causing excess elitism in the GA. It seems that the roulette selection does not behave the way we have used in our other GAs. There appears to be no weighting of individuals; rather, a threshold fitness score is collected and only individuals with a fitness score above the threshold are selected. This causes elitism, as only the most fit individuals are likely to pass this threshold. Tournament was found to be similarly troubled, with a bracket size of ⅓ of the population. This is again very elitist, as it is very large, making it very likely that only the best individuals will be selected. Before we try to further optimize the AREA GA, these selection methods should be addressed and fixed.
For roulette, we propose implementing a similar weighting scheme for individuals as is used in the PAEA GAs that gives preference to the most fit individuals, but still allows others to be selected to encourage diversity of solutions.
For crossover, we propose decreasing the bracket size. This would allow a more diverse range of individuals to be selected through “winning” smaller brackets that would have a lower probability of including top performing individuals.
4/20/23
We (Bryan and Ryan) met to work on improving the selection method functions in the AREA algorithm.
For roulette selection, we added weighting by fitness score, as is done in the other GENETIS GAs, in an effort to prevent it from disproportionately selecting the most fit individuals only to become parents. As a first test, we ran an abbreviated version of the test-loop code, printing the fitness scores of the parents selected by the roulette method. The fix appears to have passed this initial test, as the fitness scores showed more variety and in the case of roulette crossover two unique parents seemed to be selected.
For tournament selection, the only change necessary appeared to be decreasing the bracket size of the tournament(s), which in turn encourages other individuals besides the most fit to be selected as parents, therefore increasing diversity of solutions. We ran out of time to test this change fully, so we will pick up here in our next meeting.
Looking further ahead, we will plan to work on refactoring the AREA algorithm to use the same functions as the other GENETIS GAs wherever possible. In the meantime, this working version can still be used for feature development so that progress is not halted.
5/11/2023
Upon looking at the results of the recent optimizations, our best runs are still taking over 40 generations to reach an optimized score. Given the speed of AraSim, this is still too slow to be considered optimal. Comparing this test loop run to the ones run on by the broader GENETIS GA, our best runs for the AREA GA are worse than the worst runs from those optimizations. As such, we have decided to pause optimizing the AREA GA in favor of adding it to the broader GENETIS one. As of today, we have finished a rough version of the generating function necessary to create new individuals. We have also started progress on the constraining functions and scaling functions that will be necessary to generate the spherical harmonics used to find the gain and phase. Once these functions are complete, we should be able to transplant this GA back into our test loop for AREA and resume optimizations.
|
|
151
|
Fri Apr 8 16:06:31 2022 |
Aidan Snyder | AREA - frequency linear dependence correlation test run - 04/05/2022 |
- Run Type
- Run Date
- Run Name
- Why are we doing this run?
- to see if the linear dependence on frequency was properly implemented
- What is different about this run from the last?
- Under closer examination, the previous run of AREA appeared to not linearly depend on frequency, so we recompiled the GA in order to fix this
- Symmetric, asymmetric, linear, nonlinear?
- Number of individuals (NPOP)
- Number of neutrinos
- Operators / Selection methods used (% of each)
- roulette crossover 50%
- roulette mutation 16%
- tournament crossover 18%
- tournament mutation 16%
- Are we using the database?
- Result
- There is a problem with how the GA has assigned the Veff values, resulting in many individuals being assigned zero Veff, however the linear dependence seems to have been implemented successfully
Note: We also ran a run called 20220405fahimi5run1, which was a short test run with a very low amount of individuals and neutrinos which seemed to work fine; as in the linear dependence seemed to work. |
|
Draft
|
Sun Apr 5 15:07:50 2026 |
Aidan Snyder | AREA - frequency linear dependence correlation test run - 04/05/2022 |
- Run Type
- Run Date
- Run Name
- Why are we doing this run?
- to see if the linear dependence on frequency was properly implemented
- What is different about this run from the last?
- Under closer examination, the previous run of AREA appeared to not linearly depend on frequency, so we recompiled the GA in order to fix this
- Symmetric, asymmetric, linear, nonlinear?
- Number of individuals (NPOP)
- Number of neutrinos
- Operators / Selection methods used (% of each)
- roulette crossover 50%
- roulette mutation 16%
- tournament crossover 18%
- tournament mutation 16%
- Are we using the database?
- Result
- There is a problem with how the GA has assigned the Veff values, resulting in many individuals being assigned zero Veff, however the linear dependence seems to have been implemented successfully
Note: We also ran a run called 20220405fahimi5run1, which was a short test run with a very low amount of individuals and neutrinos which seemed to work fine; as in the linear dependence seemed to work. |
|
146
|
Fri Feb 11 16:34:54 2022 |
Aidan Snyder | AREA - frequency linear dependence correlation test run - 01/18/2022 |
- Run Type
- Run Date
- Run Name
- Why are we doing this run?
- to see if the linear dependence on frequency was properly implemented
- What is different about this run from the last?
- attempted to evolve gain pattern as a linear function of frequency
- Symmetric, asymmetric, linear, nonlinear?
- Number of individuals (NPOP)
- Operators / Selection methods used (% of each)
- roulette crossover 50%
- roulette mutation 16%
- tournament crossover 18%
- tournament mutation 16%
- Are we using the database?
|
| Attachment 1: Screenshot_2022-02-11_164514.png
| 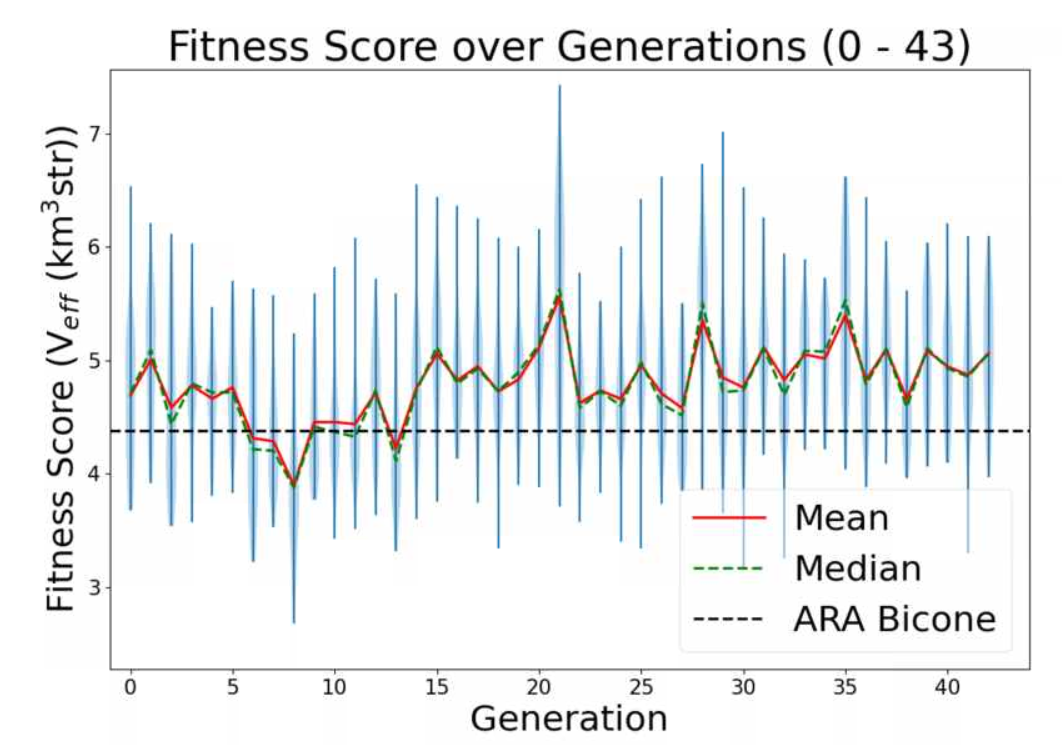 |
| Attachment 2: Screenshot_2022-02-11_164136.png
| 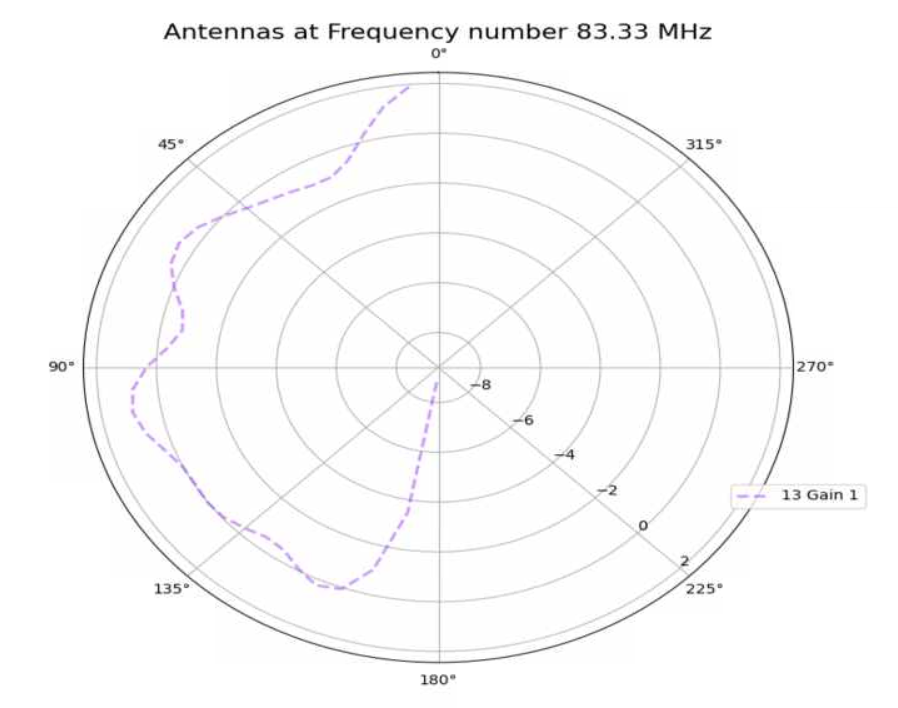 |
| Attachment 3: Screenshot_2022-02-11_164252.png
| 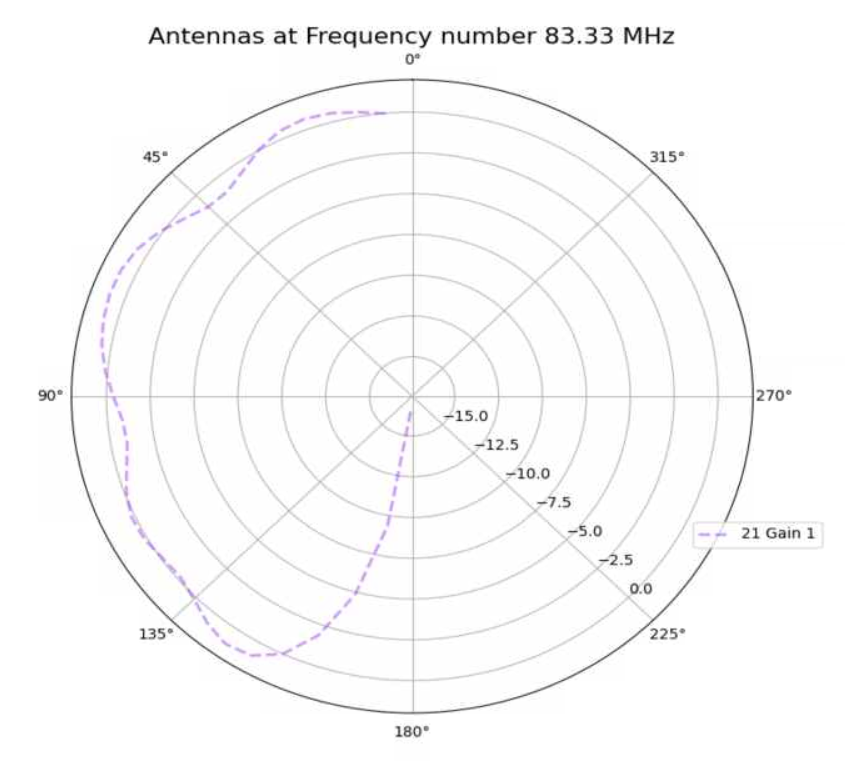 |
| Attachment 4: Screenshot_2022-02-11_164406.png
| 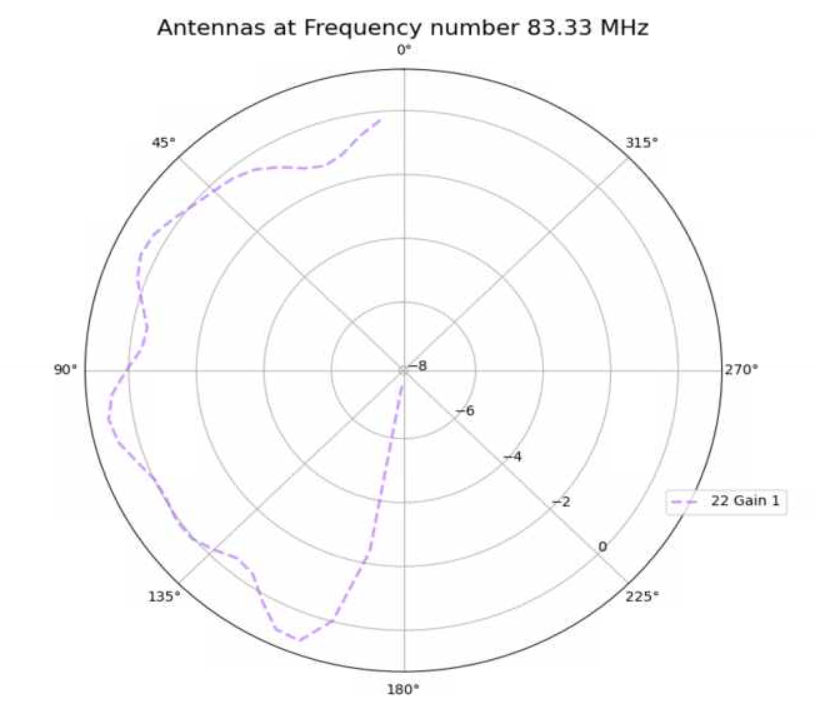 |
|
153
|
Tue Apr 19 17:46:07 2022 |
Aidan Snyder | AREA - Short run testing stringReplacement2.py - 04/19/2022 |
- Run Type
- Run Date
- Run Name
- Why are we doing this run?
- Check to see if stringReplacement2.py changes were successfull
- What is different about this run from the last?
- added below code in order to create a fitness file inside of each generation to display the fitness scores for just that generation, as opposed to just having one big file full of all the generations.
f2 = open(source + "/" + "gen_{}".format(gen) + "/fitnessFile_gen_{}".format(gen) + ".txt", 'a')
f2.write(','.join(mean_Veff_array) + "\n")
f2.close()
- Symmetric, asymmetric, linear, nonlinear?
- Number of individuals (NPOP)
- Operators / Selection methods used (% of each)
- roulette crossover 50%
- roulette mutation 16%
- tournament crossover 18%
- tournament mutation 16%
- Are we using the database?
|
|