ID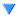 |
Date |
Author |
Subject |
|
151
|
Fri Apr 8 16:06:31 2022 |
Aidan Snyder | AREA - frequency linear dependence correlation test run - 04/05/2022 |
- Run Type
- Run Date
- Run Name
- Why are we doing this run?
- to see if the linear dependence on frequency was properly implemented
- What is different about this run from the last?
- Under closer examination, the previous run of AREA appeared to not linearly depend on frequency, so we recompiled the GA in order to fix this
- Symmetric, asymmetric, linear, nonlinear?
- Number of individuals (NPOP)
- Number of neutrinos
- Operators / Selection methods used (% of each)
- roulette crossover 50%
- roulette mutation 16%
- tournament crossover 18%
- tournament mutation 16%
- Are we using the database?
- Result
- There is a problem with how the GA has assigned the Veff values, resulting in many individuals being assigned zero Veff, however the linear dependence seems to have been implemented successfully
Note: We also ran a run called 20220405fahimi5run1, which was a short test run with a very low amount of individuals and neutrinos which seemed to work fine; as in the linear dependence seemed to work. |
|
150
|
Fri Apr 1 16:35:50 2022 |
Ryan Debolt | Population test. | https://docs.google.com/spreadsheets/d/1vvcmjByKfcns0-tbAjtePB8ZVGsXAKxXCfbMc99weMI/edit?usp=sharing Here is the spreadsheet link for the population test. |
|
149
|
Fri Mar 25 17:32:25 2022 |
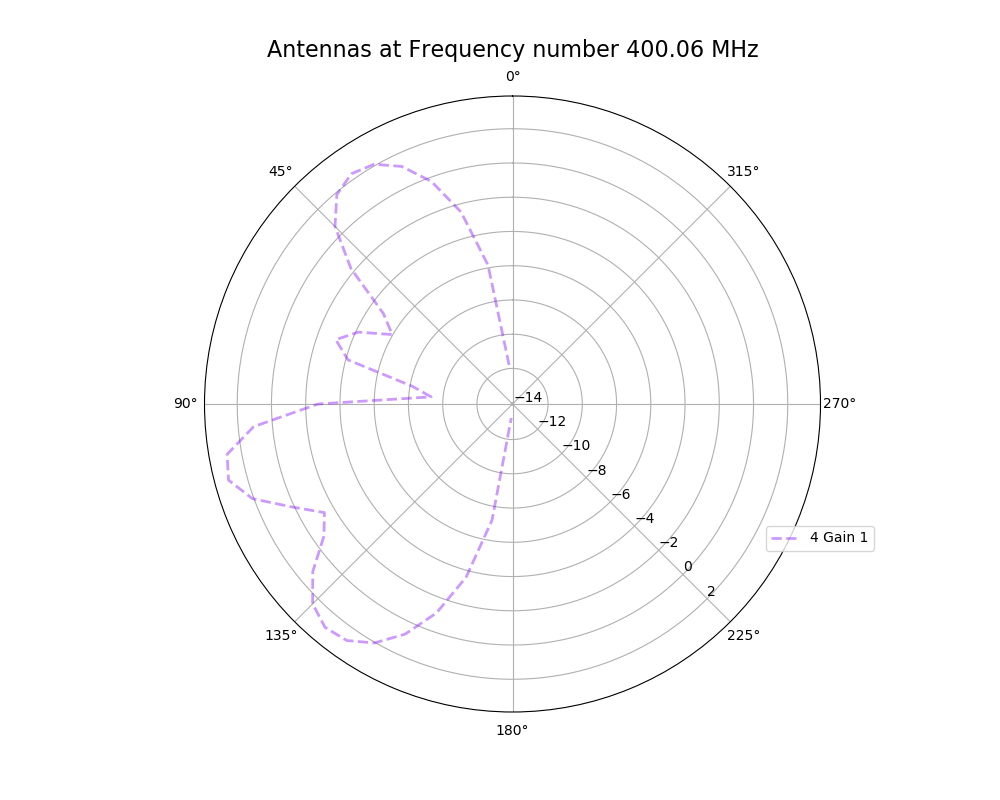
Alex M | XFdtd Step Size Investigation | The reviewer for the paper we recently submitted mentioned that our step sizes at which we are measuring the gain in XFdtd may be too large. They pointed out that there appear to be "lobes" at 400 MHz. I ran the antenna we presented in the paper through XFdtd using a step size of 5 degrees (what we've been doing) and a step size of 1 degree (the reveiwer's recommendation). Attached are antenna responses for these two different step sizes at 200 MHz and 400 MHz. Qualitatively, there are noticeable differences in the "jaggedness" of the 400 MHz plot and how extreme the minima and maxima appear, but the basic shape is preserved. We can try to do a more quantitative analysis (ex: run these through AraSim), but doing so may be more time intensive than necessary considering that AraSim may require 5 degree steps and that adding these images to an appendix may be sufficient anyway. |
| Attachment 1: polar_plot_400.06_step_5.png
|  |
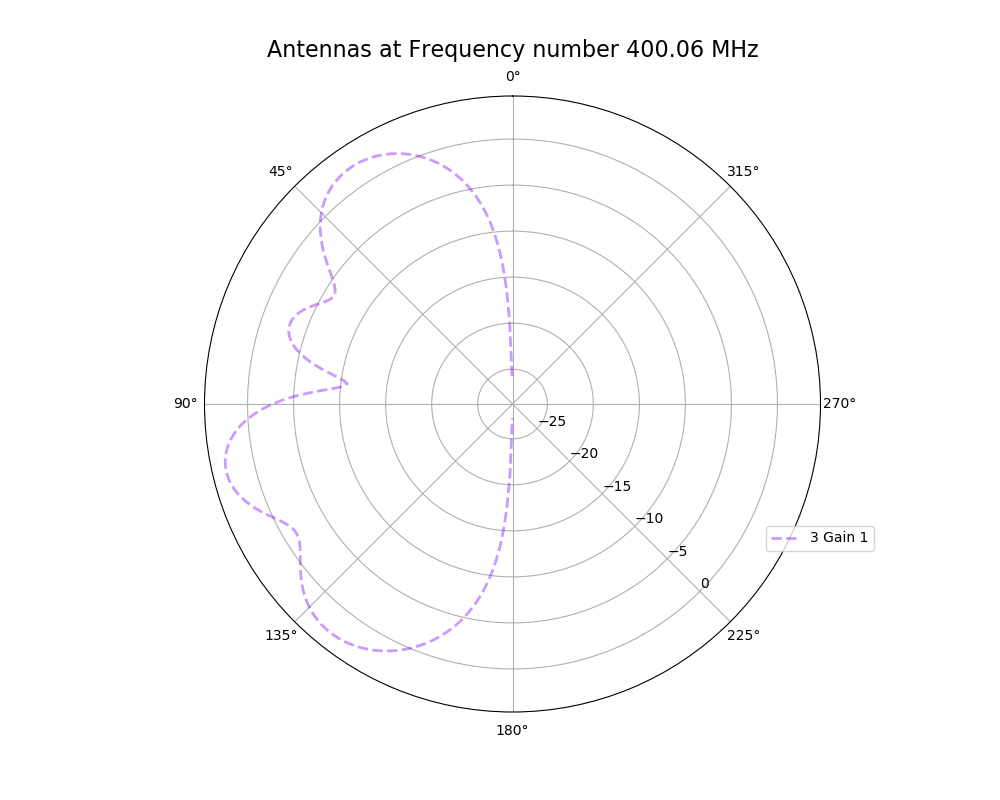
| Attachment 2: polar_plot_400.06_step_1.png
|  |
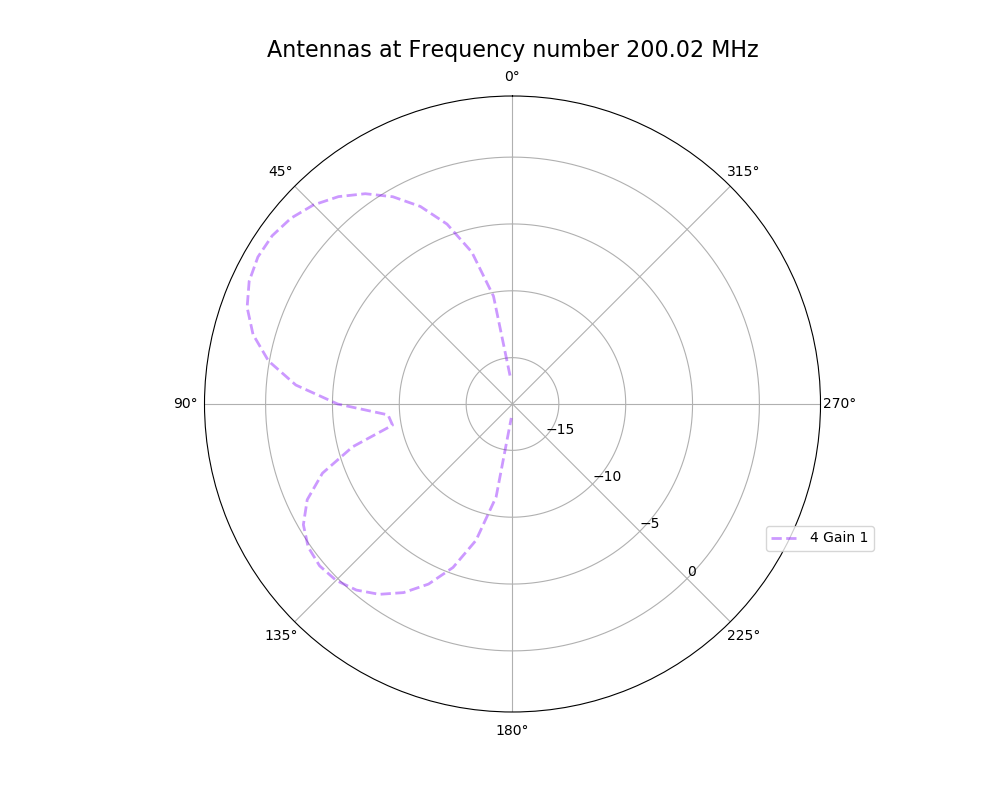
| Attachment 3: polar_plot_200.02_step_5.png
|  |
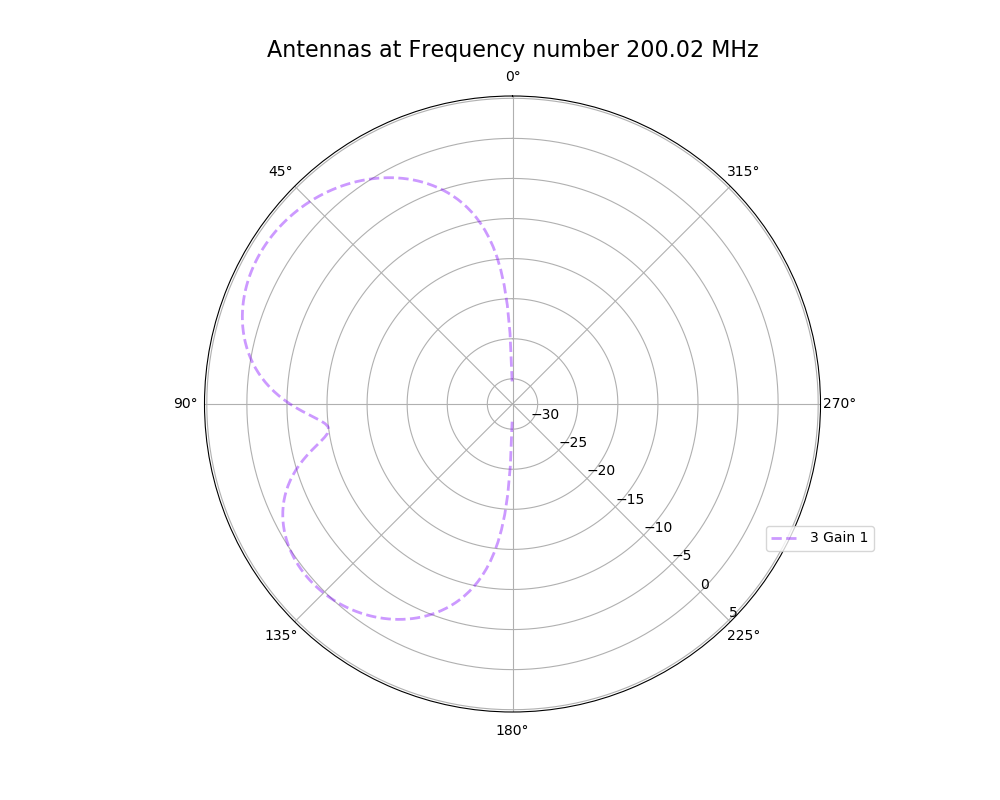
| Attachment 4: polar_plot_200.02_step_1.png
|  |
|
148
|
Fri Mar 25 17:14:01 2022 |
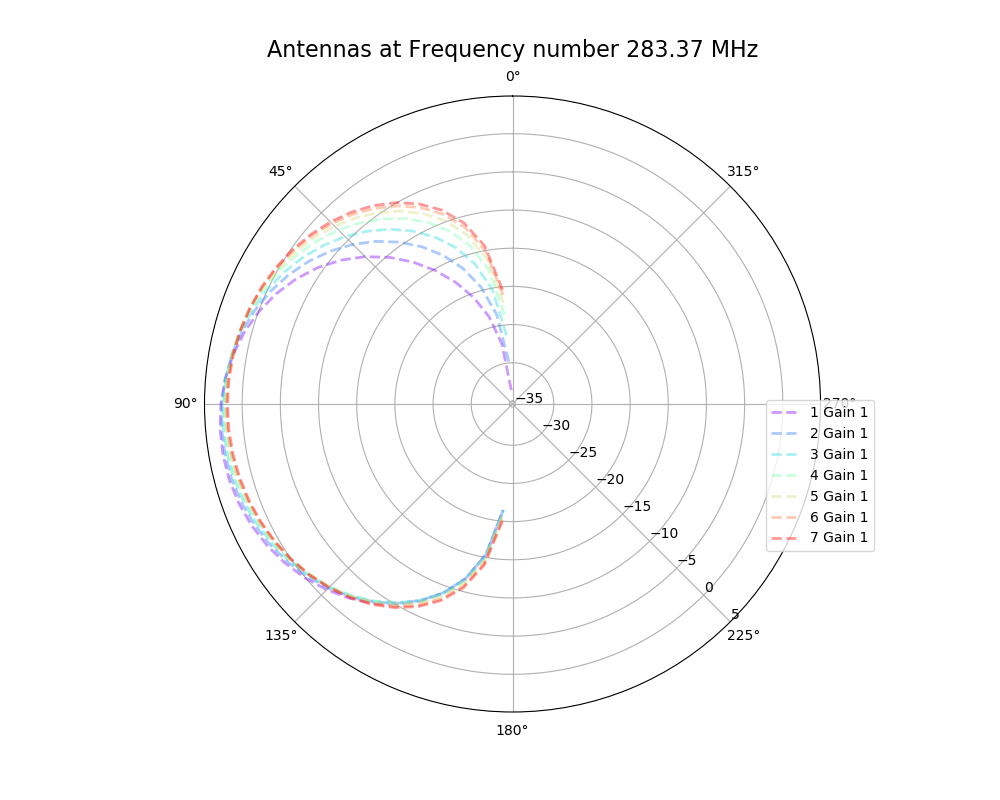
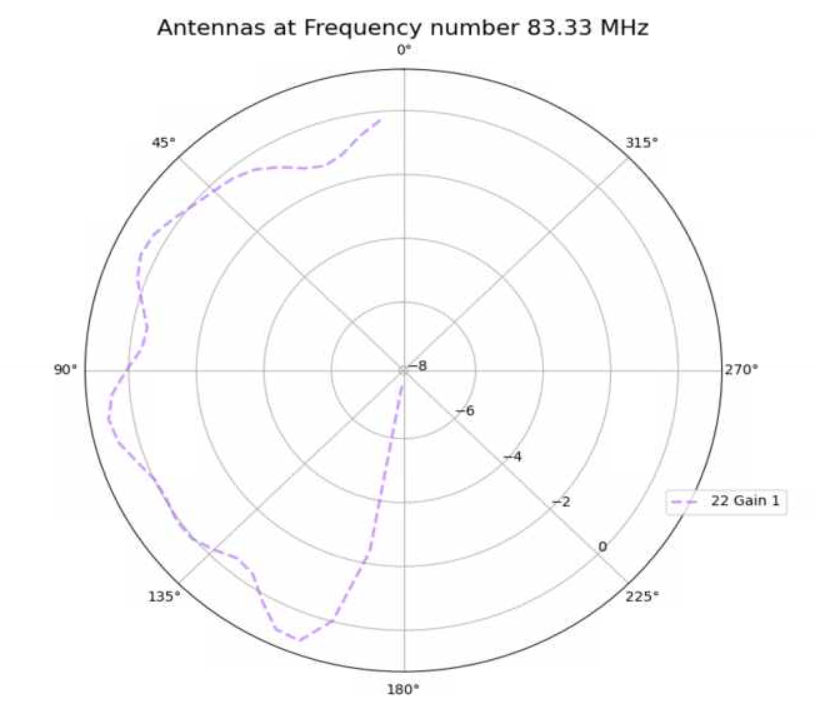
Alex M | Antenna Minimum Length Investigation | The reviewer of the paper we are trying to publish (as well as other external colleagues we showed the paper to) asked about our minimum length we implemented in the loop. Currently, we cut off the length at a minimum of 37.5 cm for each cone (minimum of 75 cm for the full cone). There is an ELOG post from Amy that presented the reasoning for this ( http://radiorm.physics.ohio-state.edu/elog/GENETIS/81 ). We initially thought that we needed a minimum because the antennas were evolving to be small (around and lower than the minimum). We thought that XFdtd was being inaccurate at lengths lower than 37.5 cm, so we set that as a minimum. To see if we can replicate any strange behavior from XFdtd, I simulated antennas at and below our minimum length and generated plots of their antennas responses. The genes are listed below, corresponding to the patterns in the plot. The results do not look unusual (that is, not dissimilar to the other bicones we have generated, including the ARA bicone) and seem to show an improved sensitivity in the upward direction for shorter bicones. The equation we used to arrive at this minimum (f = c/(4L)) might be indicative of a maximum length rather than a minimum.
| Inner radius |
Length |
Quadratic |
Linear |
2.4892,37.5,-0.00142604,0.032832
4.64522,37.5,0.00153863,-0.14004
2.4892,34.5,-0.00142604,0.032832
4.64522,34.5,0.00153863,-0.14004
2.4892,31.5,-0.00142604,0.032832
4.64522,31.5,0.00153863,-0.14004
2.4892,27.5,-0.00142604,0.032832
4.64522,27.5,0.00153863,-0.14004
2.4892,23.5,-0.00142604,0.032832
4.64522,23.5,0.00153863,-0.14004
2.4892,19.5,-0.00142604,0.032832
4.64522,19.5,0.00153863,-0.14004
2.4892,15.5,-0.00142604,0.032832
4.64522,15.5,0.00153863,-0.14004
|
| Attachment 1: polar_plot_283.37.png
|  |
|
147
|
Thu Feb 24 20:08:46 2022 |
Ryan Debolt | Parents | |
| Attachment 1: Parent_tracking_example.pdf
|
|
146
|
Fri Feb 11 16:34:54 2022 |
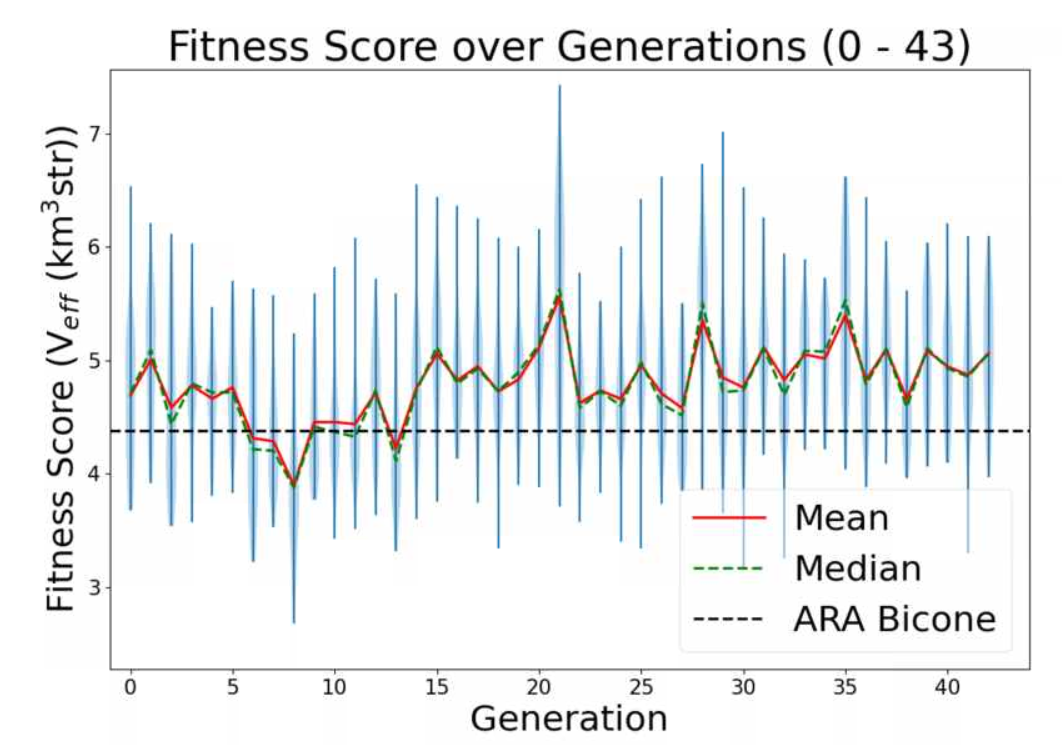
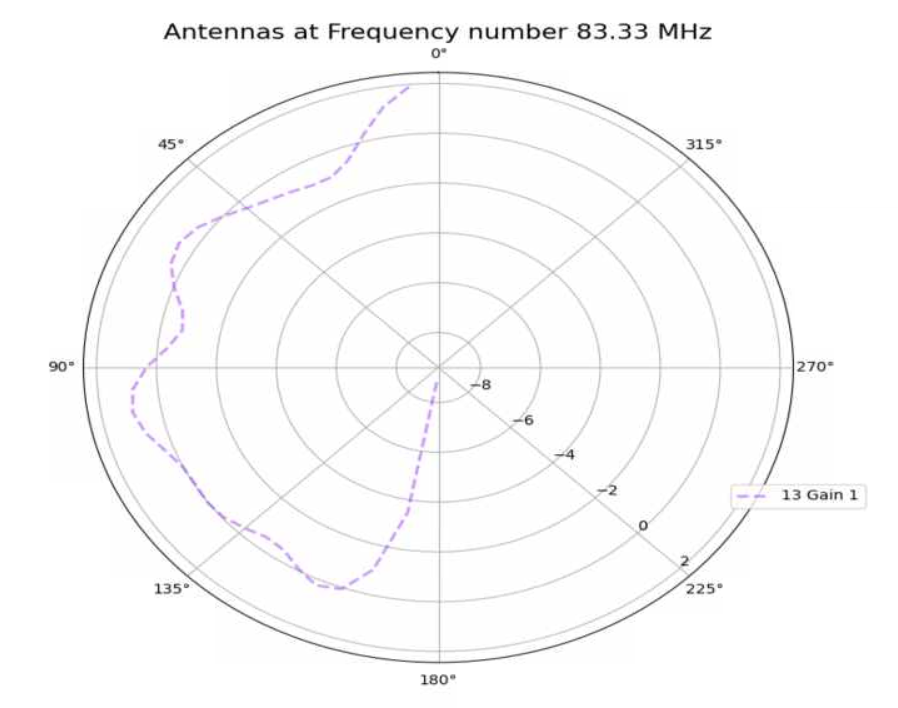
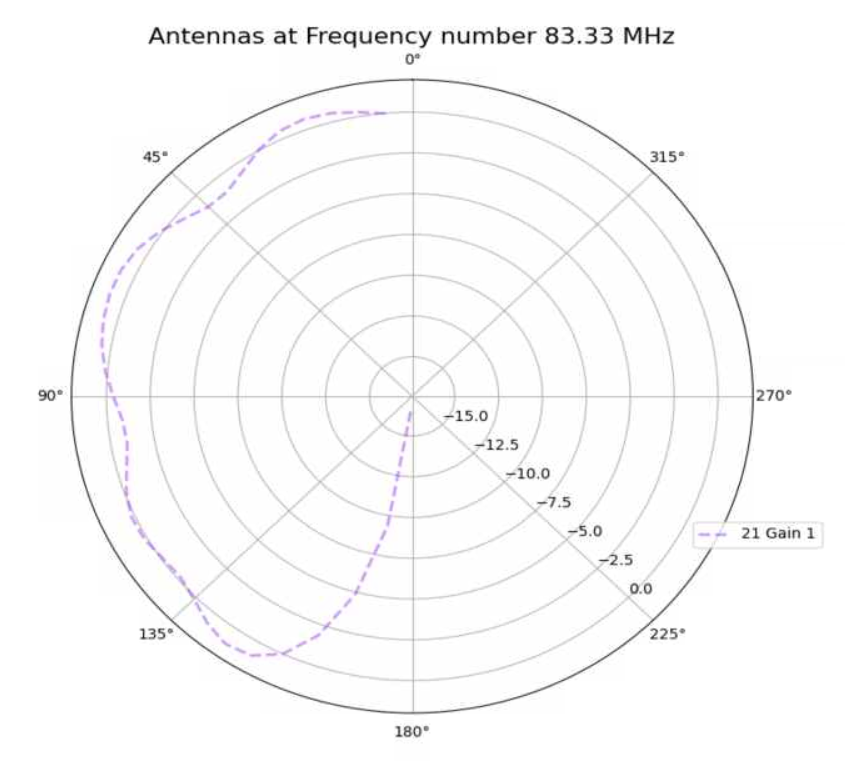
Aidan Snyder | AREA - frequency linear dependence correlation test run - 01/18/2022 |
- Run Type
- Run Date
- Run Name
- Why are we doing this run?
- to see if the linear dependence on frequency was properly implemented
- What is different about this run from the last?
- attempted to evolve gain pattern as a linear function of frequency
- Symmetric, asymmetric, linear, nonlinear?
- Number of individuals (NPOP)
- Operators / Selection methods used (% of each)
- roulette crossover 50%
- roulette mutation 16%
- tournament crossover 18%
- tournament mutation 16%
- Are we using the database?
|
| Attachment 1: Screenshot_2022-02-11_164514.png
|  |
| Attachment 2: Screenshot_2022-02-11_164136.png
|  |
| Attachment 3: Screenshot_2022-02-11_164252.png
|  |
| Attachment 4: Screenshot_2022-02-11_164406.png
|  |
|
145
|
Fri Feb 11 16:09:24 2022 |
Ryan Debolt | Parents.csv | Below is an example of our Parents.csv file written by the GA. This file tracks the parents of the individuals of the current generation. The columns and their contained information are as follows:
Current Gen:
The numbered individual of the current generation.
Parent 1:
The number of the first parent of this individual as read from the previous generation.
Parent 2:
The number of the first parent of this individual as read from the previous generation.
Operator:
The genetic operator that created the individual in that row. |
| Attachment 1: Parents.csv
|
Location of individuals used to make this generation:
Current Gen, Parent 1, Parent 2, Operator
1, 15, NA , Reproduction
2, 22, NA , Reproduction
3, 35, NA , Reproduction
4, 34, 39, Crossover
5, 34, 39, Crossover
6, 35, 29, Crossover
7, 35, 29, Crossover
8, 37, 5, Crossover
9, 37, 5, Crossover
10, 28, 12, Crossover
11, 28, 12, Crossover
12, 5, 22, Crossover
13, 5, 22, Crossover
14, 18, 23, Crossover
15, 18, 23, Crossover
16, 3, 33, Crossover
17, 3, 33, Crossover
18, 28, 14, Crossover
19, 28, 14, Crossover
20, 2, 17, Crossover
21, 2, 17, Crossover
22, 23, 22, Crossover
23, 23, 22, Crossover
24, 13, 15, Crossover
25, 13, 15, Crossover
26, 31, 35, Crossover
27, 31, 35, Crossover
28, 42, 13, Crossover
29, 42, 13, Crossover
30, 17, 6, Crossover
31, 17, 6, Crossover
32, 1, 5, Crossover
33, 1, 5, Crossover
34, 29, 38, Crossover
35, 29, 38, Crossover
36, 19, 10, Crossover
37, 19, 10, Crossover
38, 9, 38, Crossover
39, 9, 38, Crossover
40, NA, NA, Immigration
41, NA, NA, Immigration
42, NA, NA, Immigration
43, NA, NA, Immigration
44, NA, NA, Immigration
45, NA, NA, Immigration
46, NA, NA, Immigration
47, NA, NA, Immigration
48, NA, NA, Immigration
49, NA, NA, Immigration
50, NA, NA, Immigration
|
|
144
|
Tue Feb 8 15:56:42 2022 |
Machtay | Rank Test Run | I fixed a bug in the loop, so we started another rank test run. Run details in the attached file.
The bug was searching for the generationDNA.csv file in the wrong place, meaning that it wasn't able to copy it to the run directory. That meant we didn't have a record of the generation data in the usual format. I don't think that this explains the flatness, since the the generationDNA.csv file was still created every generation correctly, so the GA knew where it was. But this test now corrects that problem and tests the usage of the rank selection method. |
| Attachment 1: run_details.txt
|
####### VARIABLES: LINES TO CHECK OVER WHEN STARTING A NEW RUN ###############################################################################################
RunName='2022_02_08_Rank_Test' ## This is the name of the run. You need to make a unique name each time you run.
TotalGens=100 ## number of generations (after initial) to run through
NPOP=50 ## number of individuals per generation; please keep this value below 99
Seeds=10 ## This is how many AraSim jobs will run for each individual## the number frequencies being iterated over in XF (Currectly only affects the output.xmacro loop)
FREQ=60 ## the number frequencies being iterated over in XF (Currectly only affects the output.xmacro loop)
NNT=30000 ## Number of Neutrinos Thrown in AraSim
exp=18 ## exponent of the energy for the neutrinos in AraSim
ScaleFactor=1.0 ## ScaleFactor used when punishing fitness scores of antennae larger than the drilling holes
GeoFactor=1 ## This is the number by which we are scaling DOWN our antennas. This is passed to many files
num_keys=4 ## how many XF keys we are letting this run use
database_flag=0 ## 0 if not using the database, 1 if using the database
## These next 3 define the symmetry of the cones.
RADIUS=1 ## If 1, radius is asymmetric. If 0, radius is symmetric
LENGTH=1 ## If 1, length is asymmetric. If 0, length is symmetric
ANGLE=1 ## If 1, angle is asymmetric. If 0, angle is symmetric
CURVED=1 ## If 1, evolve curved sides. If 0, sides are straight
A=1 ## If 1, A is asymmetric
B=1 ## If 1, B is asymmetric
SEPARATION=0 ## If 1, separation evolves. If 0, separation is constant
NSECTIONS=2 ## The number of chromosomes
DEBUG_MODE=0 ## 1 for testing (ex: send specific seeds), 0 for real runs
## These next variables are the values passed to the GA
REPRODUCTION=3 ## Number (not fraction!) of individuals formed through reproduction
CROSSOVER=36 ## Number (not fraction!) of individuals formed through crossover
MUTATION=1 ## Probability of mutation (divided by 100)
SIGMA=5 ## Standard deviation for the mutation operation (divided by 100)
ROULETTE=0 ## Percent of individuals selected through roulette (divided by 10)
TOURNAMENT=1 ## Percent of individuals selected through tournament (divided by 10)
RANK=9 ## Percent of individuals selected through rank (divided by 10)
ELITE=0 ## Elite function on/off (1/0)
#####################################################################################################################################################
|
|
143
|
Fri Feb 4 17:59:41 2022 |
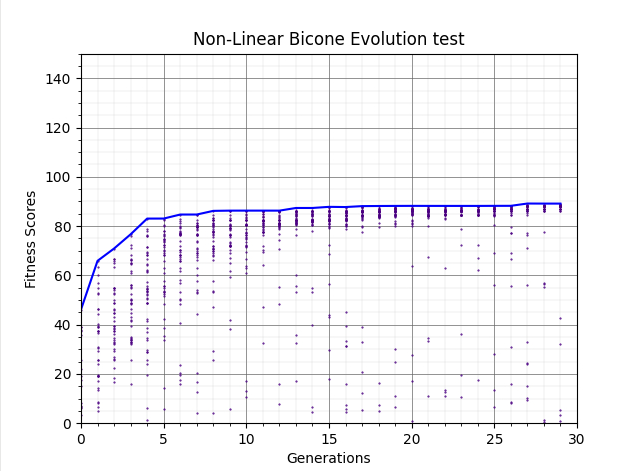
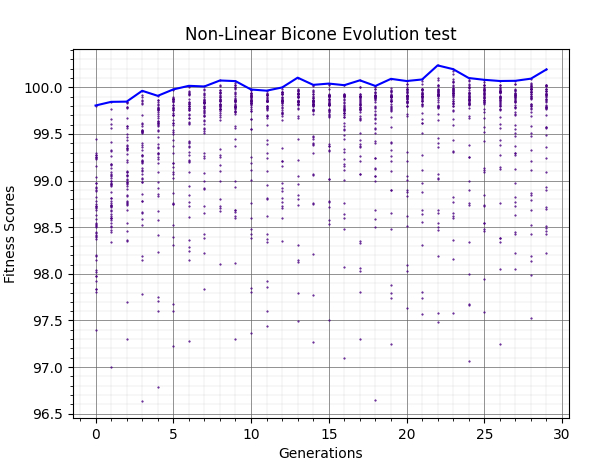
Ryan Debolt | GA Updates | The following plots are ittereations if the test loop that add increasing improvements to the GA.
The first plot shows the GA's behavoir unaltered from our previous runs (80% roulette, 20% tournament elite selection on).
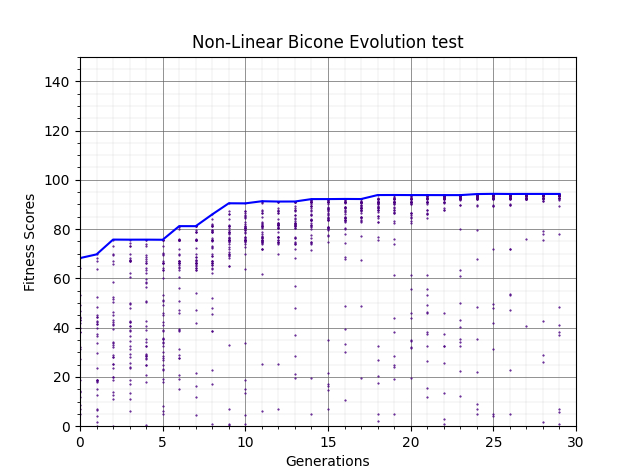
The second plots shows when we use 90% rank selection and 10% tournament, elite selection off.
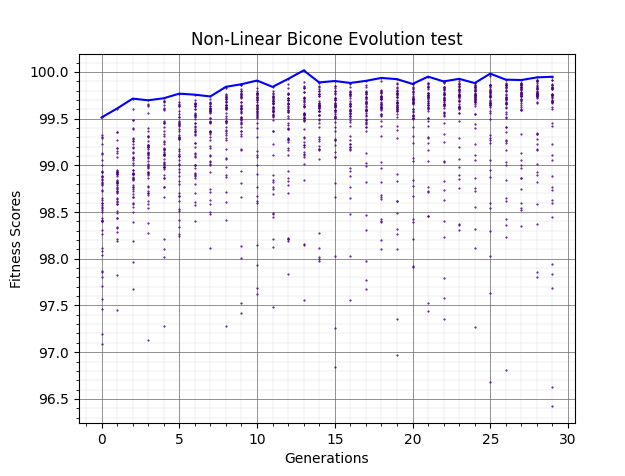
Plot 3 shows when we add an offset to restrict the values of the fitness function to be more within the range of the main loop.
Plot 4 shows when we add a gaussean mutation function that is applied to crossover individuals (rate and gaussean width chosen by guess).
The following are papers I have looked at while modifing the GA (not nessisaraly recently).
https://pdfs.semanticscholar.org/5733/418cbf21dedc9e5c04351ded4a989f1ff67e.pd
https://www.sciencedirect.com/science/article/abs/pii/0165607493902157
https://www.scientific.net/AMM.340.727
http://citeseerx.ist.psu.edu/viewdoc/download?doi=10.1.1.28.1400&rep=rep1&type=pdf
https://arxiv.org/pdf/2010.04340.pdf
https://d1wqtxts1xzle7.cloudfront.net/30694440/10.1.1.34.9722.pdf?1361979690=&response-content-disposition=inline%3B+filename%3DUsing_genetic_algorithms_with_asexual_tr.pdf&Expires=1612908683&Signature=X93Gsc47AS0xqWf1SPLjG~7sNkoXSOXfnq1GpZ2QaPrYw9x9mWwASStW2IWexo7QBzbkhzcE5tZ~CmQA1MHN-paiNFIx2ed8VNS3IhesMnotKM0mSgUZ37BCleHT9BgGkUUum8mTJBAzCUaECn6RYjm1CZpfwVPC9zwuA~DnXBST4pGlQdna22D--sHwXgX~3U3gDUSxqk8mLI0gtn~Xued3XqsTGuMUKwJ2D9UpD5yp42-3IrH6d5CZREjEfXY2geTopQ-uNkr3eOriDj0UZqSrDw5mczmod3kQrQncgd~G2Kyda4RlIs8VDzQs~BGgszHJhSDAuKDrXr8P--9tVg__&Key-Pair-Id=APKAJLOHF5GGSLRBV4ZA
http://citeseerx.ist.psu.edu/viewdoc/download?doi=10.1.1.438.7389&rep=rep1&type=pdf
https://arxiv.org/pdf/2102.01211.pdf |
| Attachment 1: Original_Params.PNG
|  |
| Attachment 2: Rank_params.PNG
|  |
| Attachment 3: Range_restriction.PNG
|  |
| Attachment 4: Mutation.PNG
|  |
|
142
|
Fri Feb 4 16:50:09 2022 |
Ryan Debolt | Loop Run |
- Run Type
- Run Date
- Run Name
- Why are we doing this run?
- To test rank selection in main loop
- What is different about this run from the last?
- Rank Slection is being used.
- Parents.csv introduced.
- Elite is being turned off.
- Symmetric, asymmetric, linear, nonlinear (what order):
- Number of individuals (NPOP):
- Number of neutrinos thrown in AraSim (NNT):
- Operatiors used (% of each):
- 06% Reproduction
- 72% Crossver
- 22% Immigration
- 1% M_rate (unused)
- 5% sigma (unused)
- Selection methods used (% of each):
- 0% Elite
- 0% Reproduction
- 10% Tournament
- 90% Rank
- Are we using the database?
Directory: /fs/ess/PAS1960/BiconeEvolutionOSC/BiconeEvolution/current_antenna_evo_build/XF_Loop/Evolutionary_Loop/Run_Outputs/2022_02_04_Rank |
| Attachment 1: run_details.txt
|
####### VARIABLES: LINES TO CHECK OVER WHEN STARTING A NEW RUN ###############################################################################################
RunName='2022_02_04_Rank' ## This is the name of the run. You need to make a unique name each time you run.
TotalGens=100 ## number of generations (after initial) to run through
NPOP=50 ## number of individuals per generation; please keep this value below 99
Seeds=10 ## This is how many AraSim jobs will run for each individual## the number frequencies being iterated over in XF (Currectly only affects the output.xmacro loop)
FREQ=60 ## the number frequencies being iterated over in XF (Currectly only affects the output.xmacro loop)
NNT=30000 ## Number of Neutrinos Thrown in AraSim
exp=18 ## exponent of the energy for the neutrinos in AraSim
ScaleFactor=1.0 ## ScaleFactor used when punishing fitness scores of antennae larger than the drilling holes
GeoFactor=1 ## This is the number by which we are scaling DOWN our antennas. This is passed to many files
num_keys=4 ## how many XF keys we are letting this run use
database_flag=0 ## 0 if not using the database, 1 if using the database
## These next 3 define the symmetry of the cones.
RADIUS=1 ## If 1, radius is asymmetric. If 0, radius is symmetric
LENGTH=1 ## If 1, length is asymmetric. If 0, length is symmetric
ANGLE=1 ## If 1, angle is asymmetric. If 0, angle is symmetric
CURVED=1 ## If 1, evolve curved sides. If 0, sides are straight
A=1 ## If 1, A is asymmetric
B=1 ## If 1, B is asymmetric
SEPARATION=0 ## If 1, separation evolves. If 0, separation is constant
NSECTIONS=2 ## The number of chromosomes
DEBUG_MODE=0 ## 1 for testing (ex: send specific seeds), 0 for real runs
## These next variables are the values passed to the GA
REPRODUCTION=3 ## Number (not fraction!) of individuals formed through reproduction
CROSSOVER=36 ## Number (not fraction!) of individuals formed through crossover
MUTATION=1 ## Probability of mutation (divided by 100)
SIGMA=5 ## Standard deviation for the mutation operation (divided by 100)
ROULETTE=0 ## Percent of individuals selected through roulette (divided by 10)
TOURNAMENT=1 ## Percent of individuals selected through tournament (divided by 10)
RANK=9 ## Percent of individuals selected through rank (divided by 10)
ELITE=0 ## Elite function on/off (1/0)
#####################################################################################################################################################
|
|
141
|
Fri Feb 4 16:10:32 2022 |
Julie Rolla | Run Log Template | For PAEA Algorithm:
Part I: Complete as soon as the run starts
Run details: Please answer all of the questions below!
- Run Type
- Answer here whether or not it's AREA (Gain pattern evolution) or PAEA (Bicone evolution)
- Run Date
- Run Name
- Parameters evolved
- Why are we doing this run?
- Add answer here about what we are testing
- What is different about this run from the last?
- Add answer here with info on what we are doing differently from the last run
- Example: We are testing a new operator, we are changing the percentages of each selection method used, etc.
- Symmetric, asymmetric, linear, nonlinear (what order):
- Add answer here (say N/A if this is an AREA run)
- Number of individuals (NPOP):
- Number of neutrinos thrown in AraSim (NNT):
- Operatiors used (% of each):
- Selection methods used (% of each):
- Are we using the database?
- Add answer here. This can be found in the main bash script within the variables section. (Answer N/A for AREA run)
Please upload the text file with all run details before closing this!
Part II: Complete as soon as the run ends
Results: Once this run completes, please upload the plot(s) to this post as an attachment, as well as a general explanation of the results.
- Summary and comments on results
- Add thoughts on what we should do next here!
- Example: "It seems like we are seeing minimal evolution. We should take a step back and try to see why we don't see improvement. Once we trouble shoot, we can start a new run to investigate."
- Example 2: "Run looks good! Our best individual is in generation #12. Attached are the CAD drawings of that individual."
**Upload all plots, run parameter text files (file that has run settings saved), CAD models of best indivduals, etc**
For AREA Algorithm:
Part I: Complete as soon as the run starts
Run details: Please answer all of the questions below!
- Run Type
- Answer here whether or not it's AREA (Gain pattern evolution) or PAEA (Bicone evolution)
- Run Date
- Run Name
- Parameters evolved
- Why are we doing this run?
- Add answer here about what we are testing
- What is different about this run from the last?
- Add answer here with info on what we are doing differently from the last run
- Example: We are testing a new operator, we are changing the percentages of each selection method used, etc.
- Single frequency run or run with broadband frequency dependence:
- Number of individuals (NPOP):
- Number of neutrinos thrown in AraSim (NNT):
- Operatiors used (% of each):
- Selection methods used (% of each):
- Any other things to note?
Please upload the text file with all run details before closing this!
Part II: Complete as soon as the run ends
Results: Once this run completes, please upload the plot(s) to this post as an attachment, as well as a general explanation of the results.
- Summary and comments on results
- Add thoughts on what we should do next here!
- Example: "It seems like we are seeing minimal evolution. We should take a step back and try to see why we don't see improvement. Once we trouble shoot, we can start a new run to investigate."
- Example 2: "Run looks good! Our best individual is in generation #12. Attached are the CAD drawings of that individual."
**Upload all plots, run parameter text files (file that has run settings saved), gain patterns of the two best and two worst individuals, etc**
|
|
140
|
Mon Nov 8 17:27:01 2021 |
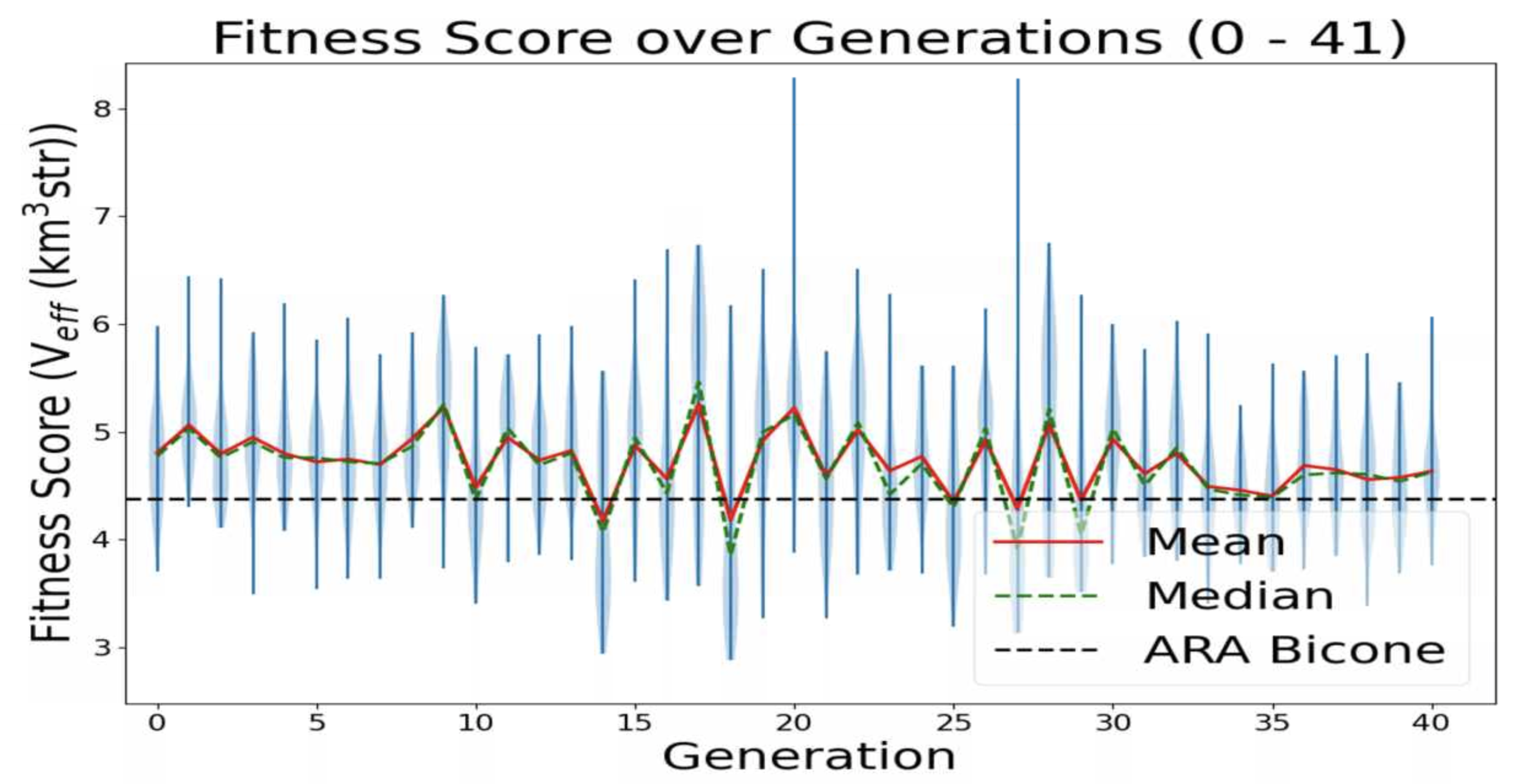
Ethan Fahimi | 07/20/2021 AREA run 3 violin plot | This is a plot made from the AREA project with full Arasim implementation with each gain pattern of each individual being fixed across all frequencies.
This run was done with 50 total individuals per generation, across 36 generations. Each individual was tested with 4 seeds of 10,000 neutrinos, for a total of 40,000 neutrinos. For each new generation, 25 individuals were created with roulette crossover, 8 with roulette mutation, 9 with tournament crossover, and 8 with tournament mutation.
individual 32 in gen 20 and individual 35 in gen 27 look promising (they have Veff > 8) |
| Attachment 1: 20211104fahimi5run2.png
|  |
|
Draft
|
Mon Nov 8 17:04:30 2021 |
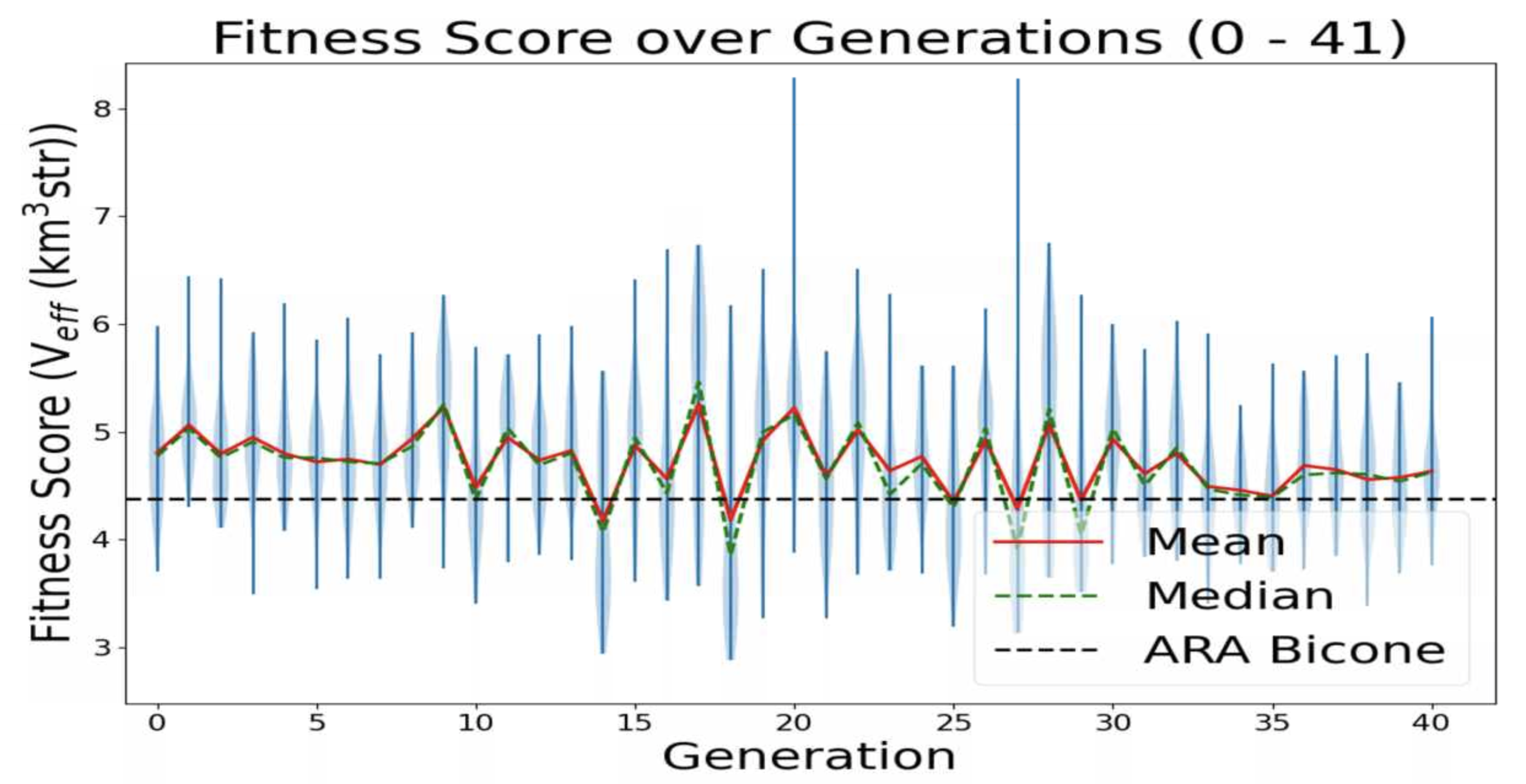
Ethan Fahimi | 11/04/2021 AREA run 2 violin plot | This is a plot made from the AREA project with full Arasim implementation with each gain pattern of each individual being fixed across all frequencies.
This run was done with 50 total individuals per generation, across 36 generations. Each individual was tested with 4 seeds of 10,000 neutrinos, for a total of 40,000 neutrinos. For each new generation, 25 individuals were created with roulette crossover, 8 with roulette mutation, 9 with tournament crossover, and 8 with tournament mutation. |
| Attachment 1: 20211104fahimi5run2.png
|  |
|
138
|
Fri Sep 17 13:41:36 2021 |
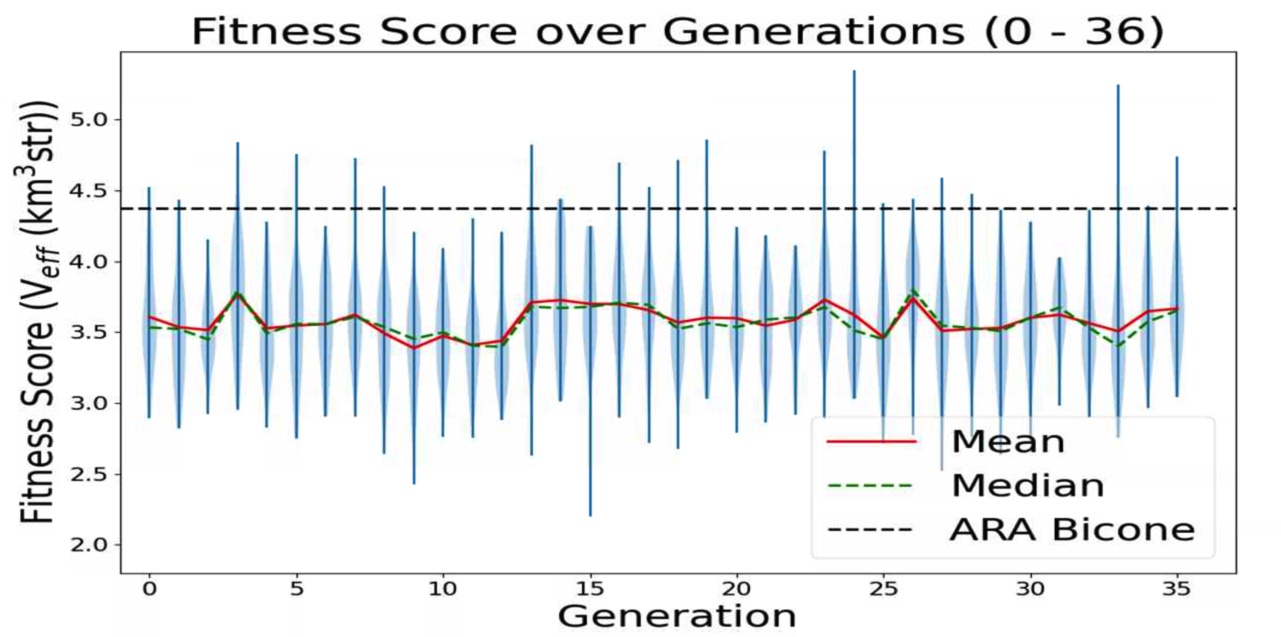
Ethan Fahimi | 07/20/2021 AREA run 3 violin plot | This is a plot made from the AREA project with full Arasim implementation. It can be seen that the Veff of any individuals is not what I would consider "good", nor is it really rising, it is quite flat. This is because in this version of AREA, the gain pattern at each frequency is generated differently than each other frequency, there is no correlation. This is known and actively being corrected. This plot is of old data and was just made for two reasons: to make sure that the violin plotting script works for AREA, to display this early form of AREA that has been adapted for full Arasim.
This run was done with 50 total individuals per generation, across 36 generations. Each individual was tested with 4 seeds of 10,000 neutrinos, for a total of 40,000 neutrinos. For each new generation, 25 individuals were created with roulette crossover, 8 with roulette mutation, 9 with tournament crossover, and 8 with tournament mutation.
This plot is further detailed in Julie Rolla's doctorate thesis. |
| Attachment 1: Image_9-13-21_at_6.35_PM.jpg
|  |
|
137
|
Wed Sep 8 16:36:33 2021 |
Alex M | MODE Workshop Presentation | We presented at a workshop put on by the MODE collaboration. MODE is a collaboration dedicated to applying Automatic Differentiation to detector design. Here's the website: https://mode-collaboration.github.io/#:~:text=MODE%20(for%20Machine%2Dlearning%20Optimized,in%20space%2C%20and%20in%20nuclear
|
| Attachment 1: GENETIS_MODE_Presentation.pptx
|
|
136
|
Fri Sep 3 14:28:55 2021 |
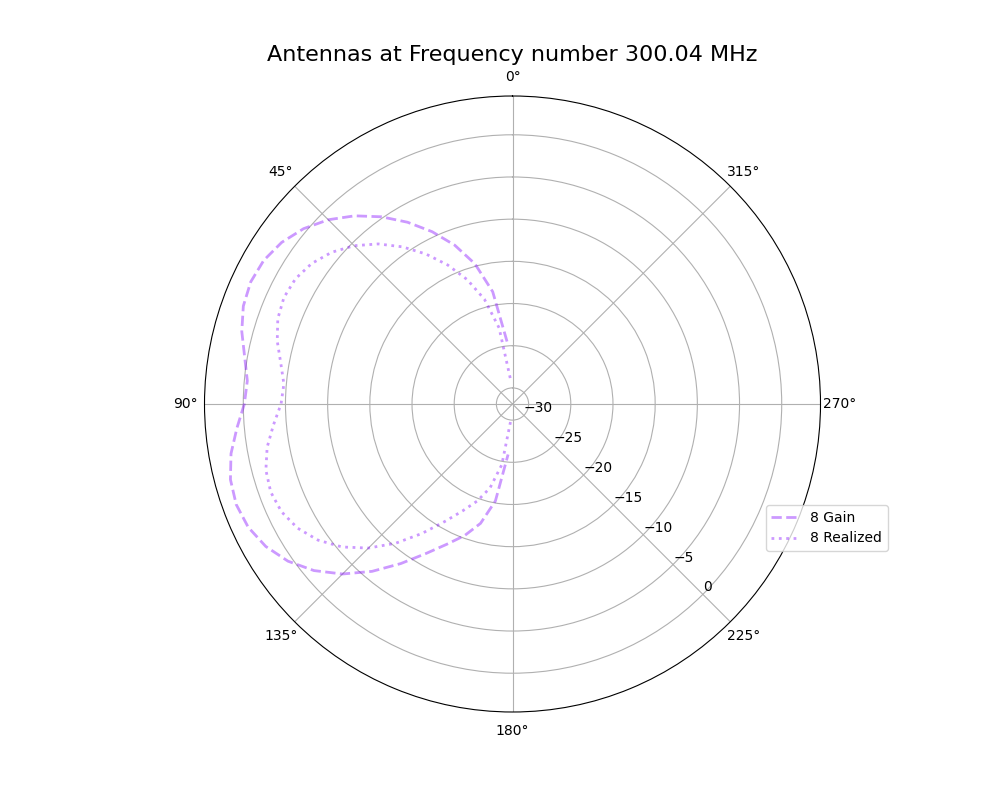
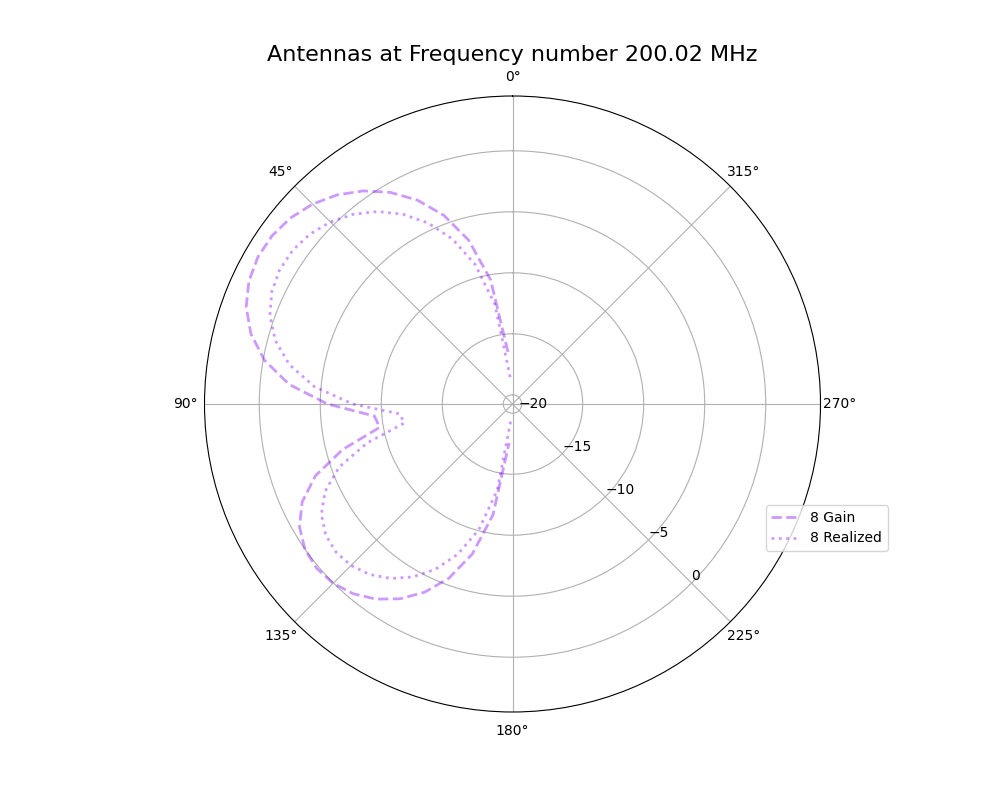
Alex M | Plots for 9/3/21 Collaboration Meeting | This ELOG post contains plots I made this week for comparing the antennas as they were evolved in the run being discussed in the upcoming paper with those same antennas when using realized gain instead of gain. These plots are preliminary, in that they should be edited before being placed in a paper (for example, VSWR is not in dBi).
Plots:
- Gain vs Realized gain
- Polar plot showing the gain and realized gain of the best individual from the run in the paper
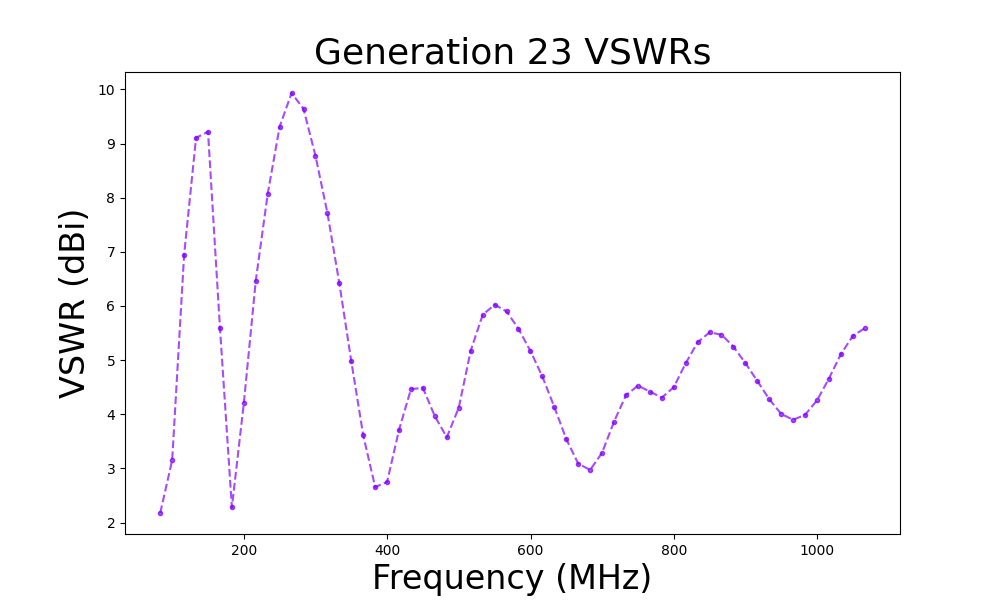
- VSWR/S11
- The VSWR of the best individual in the run over the frequency bandwidth
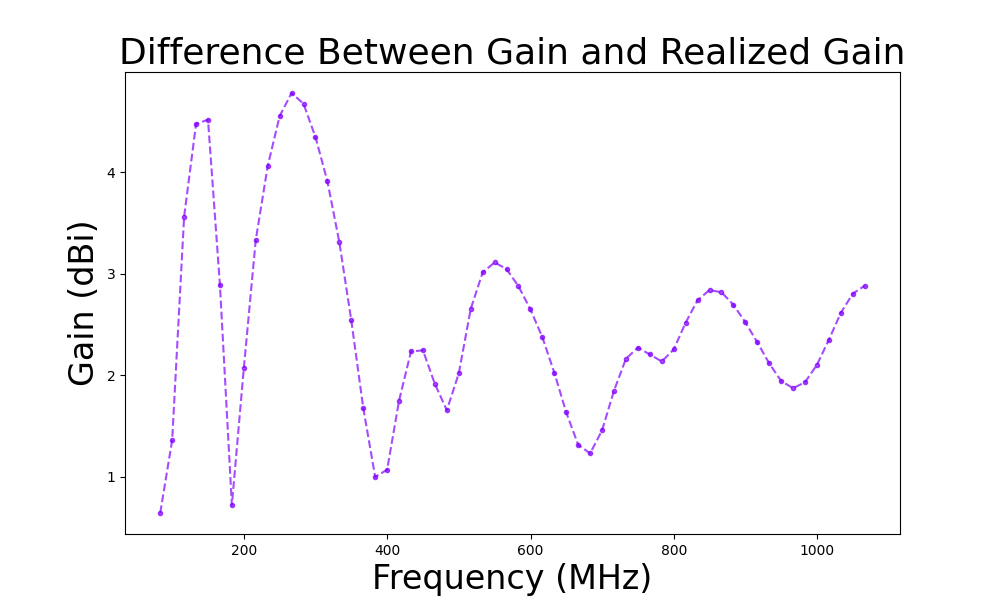
- Gain differences
- The difference between the gain and the realized gain for the best individual over the frequency bandwidth
|
| Attachment 1: polar_plot_300.04.png
|  |
| Attachment 2: polar_plot_200.02.png
|  |
| Attachment 3: VSWR_plot_1811.png
|  |
| Attachment 4: mean_gain_difference_1811.png
|  |
|
Draft
|
Fri Sep 3 14:06:39 2021 |
Alex M | Plots for 9/3/21 Collaboration Meeting | Here are plots I made for the meeting on 9/3/21. These plots represent a comparison of the gain and realized gain for the 23rd generation of the run being discussed in the upcoming paper. Here is a list of the plots
- Gain vs realized gain
- Polar plots of the best individual from generation 23
- S11/VSWR plots
- Shows the S11/VSWR over the bandwidth for the best individual
|
|
134
|
Wed Jul 14 15:45:30 2021 |
Ethan Fahimi | Wednesday Updates (7/14/2021) |
| Ethan |
Worked with Alex on fixing a few bugs with AREA. We are trying to solve an issue where the individuals are not finishing runs (around one in every four gens with 100 individuals). We believe some individuals may be too good and are then taking more than the wall time we have given them. Alex is testing this while I am working on a script that will add all the weights in the temp_{ind}.txt files. (See weightAdder.py for more) |
| |
|
| |
|
| |
|
| |
|
| |
|
| |
|
|
|
133
|
Wed Jun 23 16:34:10 2021 |
Ethan Fahimi | Daily Update 6/23/2021 |
| Name |
Progress |
Plans |
| Alex M |
|
|
| Lydon |
|
|
| Ryan |
|
|
| Ben |
|
|
| Ethan |
With Alex M's help, managed to get AREA working, plotted results. |
The results look relatively flat, possibly the GA is unoptimized, may need work. |
| Parker |
|
|
| Elliot |
|
|
| Leo |
|
|
| Evelyn |
|
|
|
|
132
|
Tue Apr 6 18:00:23 2021 |
Julie Rolla | GENETIS Google Drive with Talks/Posters, Grant writings, Papers | https://drive.google.com/drive/folders/1iDamk46R2_oOLHtvsOg4jNy05mCiB7Sn?usp=sharing |
|