| ID |
Date |
Author |
Subject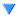 |
|
Draft
|
Mon Apr 20 01:57:35 2026 |
Julie Rolla | ONBOARDING: Tutorial Projects, Reading, Setup, etc. for New Students |
| Quote: |
|
For each new student joining the group, we ask them to onboard by completing each of the following items in the order given:
(1) Go to osc.edu and request an account
(2) Complete the following tutorials and reading. Note that the order of these does not matter, however, I would suggest "how to use Git" comes last, as it is the lease important for you to start. Finally, I advise you work on a tutorial concurrently with reading the pages assigned below from my Dissertation.
- OSC tutorials
- Because this project uses a lot of computation power, we run on the Ohio Super Computing Center's clusters. Please utilize these tutorials to learn more.
- Read Julie's Dissertation pages 1-9, 12-20 (starting at section 1.5.2), 28-37 (starting at section 2.3)
- Feel free to read chapter 3, as it has the history of this project and goes into details of our most up-to-date endeavors. This is optional and not mandatory.
- Complete the introduction to Bash project
- Complete the introduction to Python Project
- How to use Git (optional)
- Make sure to get an XFdtd license on OSC
(3) Finally, review the full manual and complete the bullets below. The full manual is Appendix A of Julie's Dissertation.
- Appendix A of my dissertation is our manual and acts as the holy grail of our research. Anything you need can be found here (from where our info exists on the web, to how to run the code).
- Once you have read this through entirely, we advise you log on to OSC and explore the code for PAEA in PAS1960. Once you have done so, complete this homework by making a copy of this google doc and filling it out on your own.
Once these are completed, the students are ready to start helping. This is all of the coding information and basic background information on evolutionary algorithms students must know to be proficient in assisting.
Other useful materials we have may include:
- Our old GENETIS manual
- This is somewhat out of date; however, it does have information on the general working on the loop.
- Julie's candidacy paper
- Complete the writing a genetic algorithm in C++ project
- This is still a work in progress! I'll complete it shortly!
- Read chapter 3 of this book (more if you have time) by Eiben and Smith, "Introduction to Evolutionary Computation" for a quick introduction. The book should be available electronically from a university library. Let us know if you can't find it there.
- Get to know XFdtd
- XFdtd manual is also attached; however, the link above (in 5) is more useful for coding a .xmacro.
Important notes:
MAC USERS: Using a GUI when connecting to OSC via SSH from your terminal.
You will need to set up X11 forwarding. Otherwise, when you go to plot via SSH, they will not display (they don't forward from OSC to your machine). If you are unable to run GUI's on OSC through your computer's terminal, follow these instructions:
- In your own system's terminal, type: sudo vi /etc/ssh/sshd_config
- Search for the following line: X11Forwarding no
- Change it to: X11 Forwarding yes. Save the file and exit.
- In your own system's terminal, type: sudo systemctl restart sshd
- For a Mac, download and install XQuartz.
- Connect to OSC using ssh -XY username@hostname.
- You can test that this works by typing 'xeye' or 'xclock' in the terminal when logged into OSC.
Now you should be able to use OSC's GUIs.
PC USERS:
As a PC user, there are a few things you need to do:
(1) Enable the Windows Subsystem for Linux on your device (https://developerinsider.co/stepwise-guide-to-enable-windows-10-subsystem-for-linux/#:~:text=To%20enable%20the%20Windows%20Subsystem,list%20here%20and%20click%20OK.)
(2) Get Bash on Ubuntu installed on your machine. You are going to need to have Bash on Ubuntu installed prior to starting these trainings; otherwise Bash commands will not be recognized.
(3) Get X11 forwarding working. Otherwise, when you go to plot via SSH, they will not display (they don't forward from OSC to your machine).
- Follow these steps here.
- Change your config file. To do so, in terminal (with Bash on Ubuntu open)
- type: sudo vi /etc/ssh/sshd_config
- Then change '#X11Forwarding no' to '#X11Forwarding yes' and save it.
- Make sure Putty and Xming are running every time before you ssh in. Use Xlaunch application to run Xming and press yes to everyrthing.
- Xming should now be running as a background task.
- Open PuTTY and open the pre-saved like you saved in PuTTY. (This will open a Bash terminal that asks for your OSC login password.)
|
|
|
9
|
Wed Feb 27 16:29:35 2019 |
Julie Rolla | New To-Do List | Here are the following things we need to work on:
- Paper (Julie and Suren)
- Get loop working-- specifically XF (Cade)
- Test GA (Suren)
- Gain pattern plots -- Make them 3D (Evelyn) & put on GitHub
- Insert LR plot & Gain plot to loop (?)
- Edit bash script
- Update Github
- AraSim plots (Julie and Max) -- see #2 and #3 here: http://radiorm.physics.ohio-state.edu/elog/GENETIS/7 for details.
- Update manual with things from Suren's last post (?)
|
|
131
|
Fri Mar 19 17:39:47 2021 |
Alex M | New Run | We began a new run using the same parameters as the previous one (see the last ELOG entry). The previous one ran for 8 generations. The difference between this run and the last one is that we have added in the polarity fix for AraSim Brian and Jorge gave us (copying the correct Report.cc and Report.h files into our AraSim version). Here are the run parameters:
Run Paramaters:
50 individuals
80/20 Roulette/Tournament
6% Reproduction, 72% Crossover, 22% Mutation
Run for 10 generations, then implement Jorge and Brian's fix for polarization in AraSim for the next run |
|
Draft
|
Thu Jun 30 13:04:48 2022 |
Ryan Debolt | Multigenerational Narrative draft 2 | This is a multigenerational tracing of our second-best individual's parents and children:
The second-best individual in this evolution was Bicone 22 from generation 40. This individual is, in fact, a fascinating case as we shall see. But to start the story of this individual we will go back to generation 38 in order to demonstrate some of the peculiarities.
In generation 38 there were two bicones of no renown, Bicone 11 and Bicone 17. Bicone 11 was a fairly average individual that was ranked 23rd with a fitness score of 4.72016. Its DNA was {*6.16084, ***79.663, 0.0015434, -0.107765} for side one , and {0.308809, 39.6742, -0.0084247, 0.40629} for side two. One day, by chance it met Bicone 17, another average bicone ranked 24th with a score of 4.71877 and DNA {**2.32499, 79.663, -0.00224948, 0.192602} for side one, and {0.308809, 39.6742, -0.0084247, 0.40629} for side two. These two individuals eventually became the parents of two antennas: Bicone 16 and Bicone 17 of generation 39.
Bicone 16 eventually grew to have been ranked 3rd in fitness score of 4.95323. It’s DNA ended up being a complete balance of its parents sharing side one with Bicone 38.11{2.32499, 79.663, -0.00224948, 0.192602} and side two with bicone 38.17 {0.308809, 39.6742, -0.0084247, 0.40629}. Bicone 16 was an individual with high aspirations and hoped to be reproduced. But alas, it was not meant to be. But bicone 16 came upon some amazing luck, it was selected with itself for crossover. This meant that bicone 16 was able to provide two identical surrogates to survive into the next generation. This is where this Bicone fulfilled its full potential.
The twin bicones were named Bicone 22 and Bicone 23 in generation 40. Being clones, they shared all their DNA with Bicone 39.16. However, due to some circumstances, they had slightly different fitness scores. Bicone 23 managed a very respectable 5.0117 fitness score and was ranked 2nd in the generation. But not to be outdone Bicone 22 managed to score a 5.17014 and ended up being the second-best performing individual of all time. Being so successful, the two bicones ended up producing 8 children between the two groups.
Bicone 23 was the first to crossover and had 2 children with its partner. These were Bicones 4 and 5. Bicone 4 was ranked 39th with a fitness score of 4.60251 and still shared the DNA of its second side with its grandparent Bicone 38.17 as well as most of its first side with Bicone 38.11 {2.32499, 79.663, -0.00213879, 0.192602} {0.308809, 39.9608, -0.0084247, 0.40629}. Bicone 5 on the other hand, was ranked 14th with a fitness score of 4.80535 and it still shared a lot of DNA with its grandparents {6.16084,53.0851,-0.00224948,0.0534469} {0.966617,39.6742,-0.0084247,0.40629}.
Bicone 22 had 6 children of its own with various other Bicones. These were; Bicone 8, ranked 11th with a fitness Score of 4.81784 and DNA {2.32499, 75.9855, -0.00224948, 0.192602} {0.966617, 39.6742, -0.00320023, 0.213833}; Bicone 9, ranked 7th with a fitness score of 4.88966 and DNA {6.10508, 79.663, -0.000594616, 0.0351901} {0.308809, 42.4246, -0.0084247, 0.40629}; Bicone 12, ranked 12th with a fitness score of 4.81705 and DNA {6.42695, 75.9855, 0.0015434, -0.107765} {0.308809, 39.6742, -0.00320023, 0.213833}; Bicone 13, ranked 2nd with a fitness score of 5.0344 and DNA {2.32499, 79.663, -0.00224948, 0.192602} {0.966617, 42.4246, -0.0084247, 0.40629}; Bicone 34 ranked 3rd with a fitness score of 4.99864 and DNA {0.66148, 73.5522, -0.000594616, 0.00582814} {0.966617, 39.6742, -0.0084247, 0.40629}; and finally Bicone 35, ranked 22nd with fitness score 4.74955 and DNA {2.32499, 79.663, -0.00224948, 0.192602} {0.308809, 42.4246, -0.0084247, 0.40629}
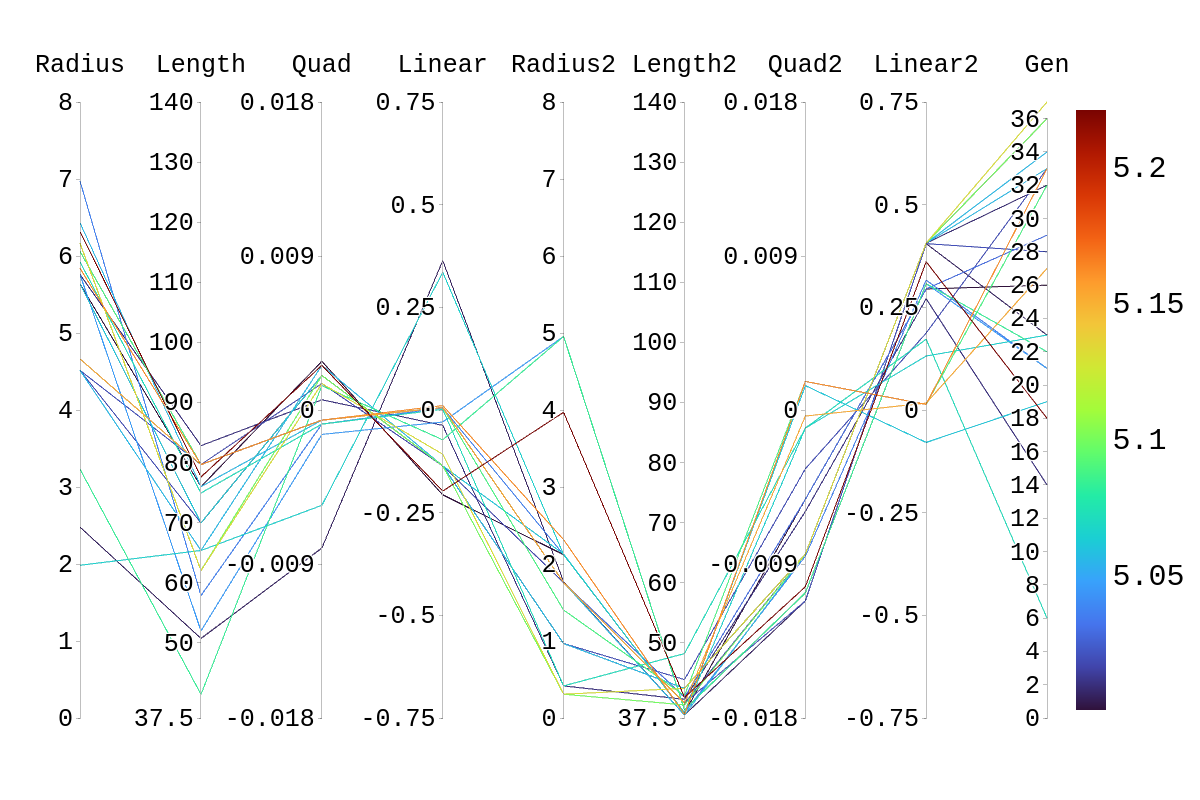
Bellow, I have attached the rainbow plot with the parameters occupied by individual 4 in gen 41 which was again ranked 39th in that generation. From this, we can see that while in its own generation it was a poor performer, overall it was upper middle of the pack. However, because of the density of other better performing antennas in this region, it is hard to distinguish which genes in this antenna are contributing the most to the drop in fitness score compared to its siblings and parents.
*Gene originating from Bicone 38.11
**Gene originating from Bicone 38.17
***Gene originating from Bicone 38.11 and 38.17 that is shared between the two. |
| Attachment 1: rainbow_plot_quadratic_rank_test_720_(drawn).png
| .png) |
|
Draft
|
Fri Jul 8 13:34:08 2022 |
Ryan Debolt | Multigenerational Narrative draft 2 | |
|
173
|
Fri Jul 8 13:35:19 2022 |
Ryan Debolt | Multigenerational Narrative draft 2 | The second-best individual in this evolution was Bicone 22 from generation 40. This individual is, in fact, a fascinating case as we shall see. But to start the story of this individual we will go back to generation 38 in order to demonstrate some of the peculiarities.
In generation 38 there were two bicones of no renown, Bicone 11 and Bicone 17. Bicone 11 was a fairly average individual that was ranked 23rd with a fitness score of 4.72016. Its DNA was {*6.16084, ***79.663, 0.0015434, -0.107765} for side one , and {0.308809, 39.6742, -0.0084247, 0.40629} for side two. One day, by chance it met Bicone 17, another average bicone ranked 24th with a score of 4.71877 and DNA {**2.32499, 79.663, -0.00224948, 0.192602} for side one, and {0.308809, 39.6742, -0.0084247, 0.40629} for side two. These two individuals eventually became the parents of two antennas: Bicone 16 and Bicone 17 of generation 39.
Bicone 16 eventually grew to have been ranked 3rd in fitness score of 4.95323. It’s DNA ended up being a complete balance of its parents sharing side one with Bicone 38.11{2.32499, 79.663, -0.00224948, 0.192602} and side two with bicone 38.17 {0.308809, 39.6742, -0.0084247, 0.40629}. Bicone 16 was an individual with high aspirations and hoped to be reproduced. But alas, it was not meant to be. But bicone 16 came upon some amazing luck, it was selected with itself for crossover. This meant that bicone 16 was able to provide two identical surrogates to survive into the next generation. This is where this Bicone fulfilled its full potential.
The twin bicones were named Bicone 22 and Bicone 23 in generation 40. Being clones, they shared all their DNA with Bicone 39.16. However, due to some circumstances, they had slightly different fitness scores. Bicone 23 managed a very respectable 5.0117 fitness score and was ranked 2nd in the generation. But not to be outdone Bicone 22 managed to score a 5.17014 and ended up being the second-best performing individual of all time. Being so successful, the two bicones ended up producing 8 children between the two groups.
Bicone 23 was the first to crossover and had 2 children with its partner Bicone 25 (4.64409). These were Bicones 4 and 5. Bicone 4 was ranked 39th with a fitness score of 4.60251 and still shared the DNA of its second side with its grandparent Bicone 38.17 as well as most of its first side with Bicone 38.11 {2.32499, 79.663, -0.00213879, 0.192602} {0.308809, 39.9608, -0.0084247, 0.40629}. Bicone 5 on the other hand, was ranked 14th with a fitness score of 4.80535 and it still shared a lot of DNA with its grandparents {6.16084,53.0851,-0.00224948,0.0534469} {0.966617,39.6742,-0.0084247,0.40629}.
Bicone 22 had 6 children of its own with various other Bicones. These were; With Bicone 8.40 (4.82423) Bicone 8, ranked 11th with a fitness Score of 4.81784 and DNA {2.32499, 75.9855, -0.00224948, 0.192602} {0.966617, 39.6742, -0.00320023, 0.213833}; Bicone 9, ranked 7th with a fitness score of 4.88966 and DNA {6.10508, 79.663, -0.000594616, 0.0351901} {0.308809, 42.4246, -0.0084247, 0.40629}; With Biocone 16.40 (5.01137) Bicone 12, ranked 12th with a fitness score of 4.81705 and DNA {6.42695, 75.9855, 0.0015434, -0.107765} {0.308809, 39.6742, -0.00320023, 0.213833}; Bicone 13, ranked 2nd with a fitness score of 5.0344 and DNA {2.32499, 79.663, -0.00224948, 0.192602} {0.966617, 42.4246, -0.0084247, 0.40629}; With Bicone 4.40 (4.66578) Bicone 34 ranked 3rd with a fitness score of 4.99864 and DNA {0.66148, 73.5522, -0.000594616, 0.00582814} {0.966617, 39.6742, -0.0084247, 0.40629}; and finally Bicone 35, ranked 22nd with fitness score 4.74955 and DNA {2.32499, 79.663, -0.00224948, 0.192602} {0.308809, 42.4246, -0.0084247, 0.40629}
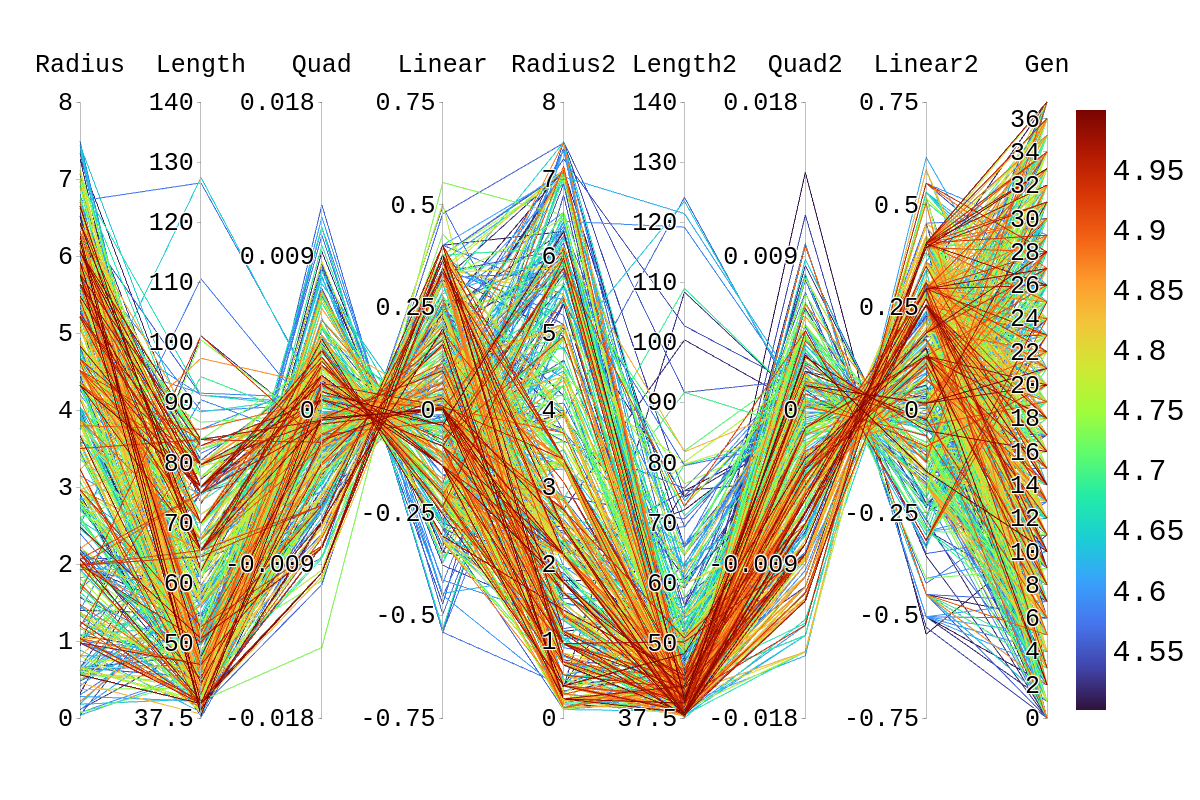
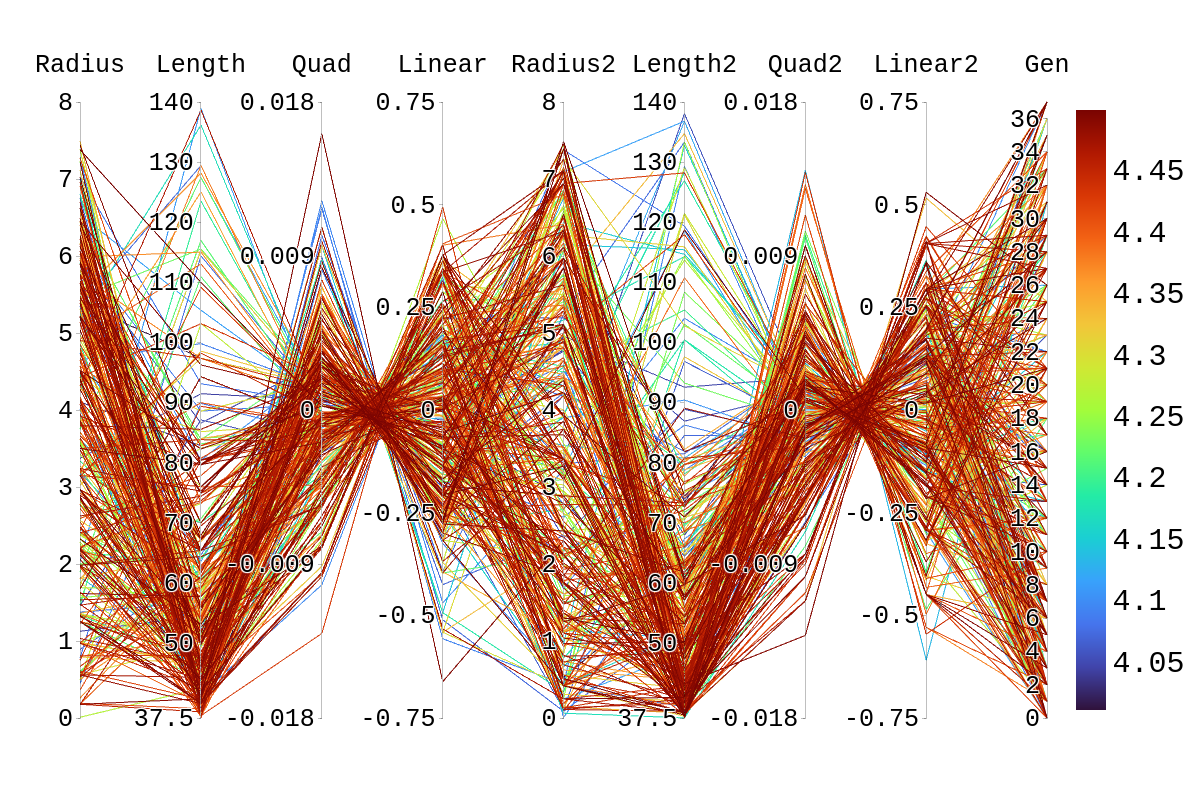
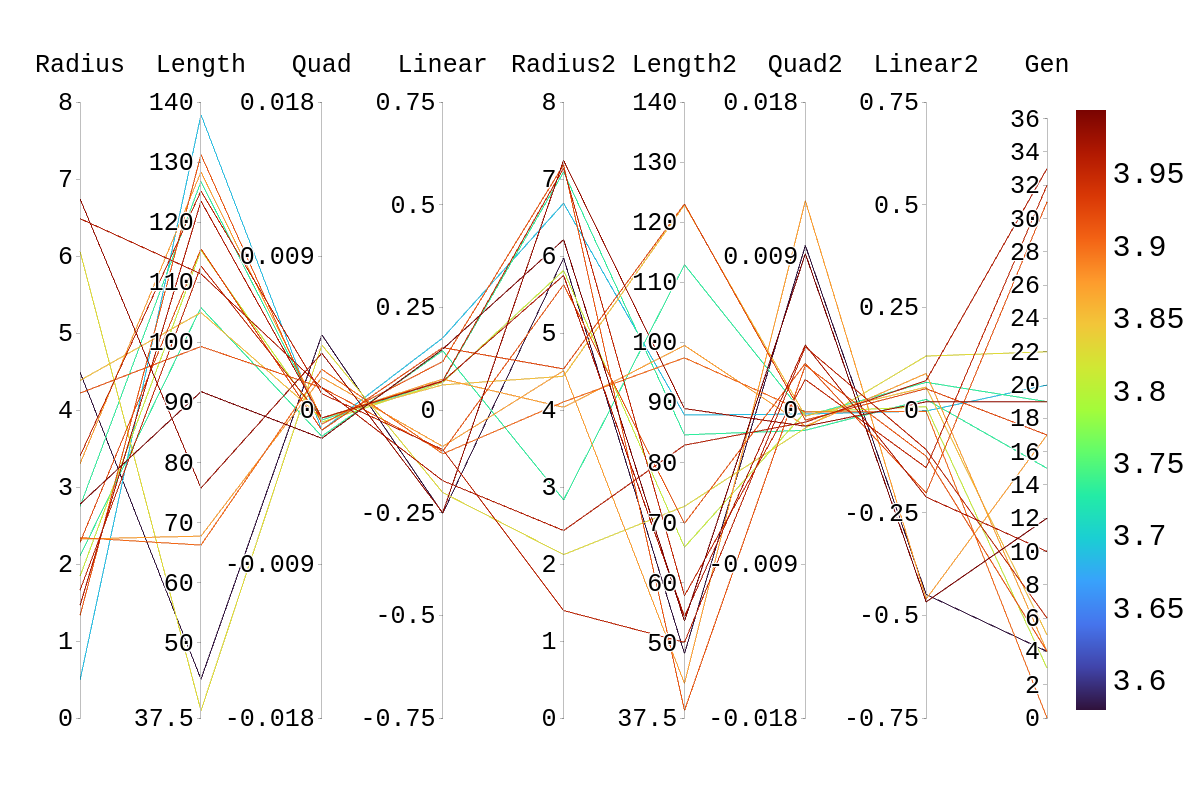
Upon inspection of the children in generation 41, you will see that none of the children surpassed their parent individual with the highest fitness score. They do however tend to have fitness scores around or above their second parent's score. Therefore, it is likely that the children's lower fitness score is due to a mismatch of genes. If we look at the rainbow plots provided, we can see that the genes of the worst-performing child are not dissimilar to those of individuals that scored higher than 5. This would imply that the fitness score found could be due to variations in Arasim similar to those seen from individuals 16.39 to 22/23.40 despite them being identical. Though it also could be possible that the graphs do not have a fine enough display to be able to clearly tell some of the genes apart.
*Gene originating from Bicone 38.11
**Gene originating from Bicone 38.17
***Gene originating from Bicone 38.11 and 38.17 that is shared between the two.
Amy: We want to nail things down in places where we still have hypotheses for what could have happened. For example, "This would imply that the fitness score found could be due to variations in Arasim similar to those seen from individuals 16.39 to 22/23.40 despite them being identical. Though it also could be possible that the graphs do not have a fine enough display to be able to clearly tell some of the genes apart." These two hypotheses can easily be distinguished by running with higher statistics and looking at the numbers rather than relying on the resolution of the display.
Amy: Also, at each stage, I was looking for the random numbers that were generated that led to the selection of the fractions of the population that were used for each type of selection and operator.
|
| Attachment 1: Rainbow_Plot_Quadratic_Rank_Test_5.png
|  |
| Attachment 2: Rainbow_Plot_Quadratic_Rank_Test_4.5.png
|  |
| Attachment 3: Rainbow_Plot_Quadratic_Rank_Test_4.png
|  |
| Attachment 4: Rainbow_Plot_Quadratic_Rank_Test_3.5.png
|  |
|
Draft
|
Thu Mar 26 02:18:17 2026 |
Ryan Debolt | Multigenerational Narrative draft 2 | This is a multigenerational tracing of our second-best individual's parents and children:
The second-best individual in this evolution was Bicone 22 from generation 40. This individual is, in fact, a fascinating case as we shall see. But to start the story of this individual we will go back to generation 38 in order to demonstrate some of the peculiarities.
In generation 38 there were two bicones of no renown, Bicone 11 and Bicone 17. Bicone 11 was a fairly average individual that was ranked 23rd with a fitness score of 4.72016. Its DNA was {*6.16084, ***79.663, 0.0015434, -0.107765} for side one , and {0.308809, 39.6742, -0.0084247, 0.40629} for side two. One day, by chance it met Bicone 17, another average bicone ranked 24th with a score of 4.71877 and DNA {**2.32499, 79.663, -0.00224948, 0.192602} for side one, and {0.308809, 39.6742, -0.0084247, 0.40629} for side two. These two individuals eventually became the parents of two antennas: Bicone 16 and Bicone 17 of generation 39.
Bicone 16 eventually grew to have been ranked 3rd in fitness score of 4.95323. It’s DNA ended up being a complete balance of its parents sharing side one with Bicone 38.11{2.32499, 79.663, -0.00224948, 0.192602} and side two with bicone 38.17 {0.308809, 39.6742, -0.0084247, 0.40629}. Bicone 16 was an individual with high aspirations and hoped to be reproduced. But alas, it was not meant to be. But bicone 16 came upon some amazing luck, it was selected with itself for crossover. This meant that bicone 16 was able to provide two identical surrogates to survive into the next generation. This is where this Bicone fulfilled its full potential.
The twin bicones were named Bicone 22 and Bicone 23 in generation 40. Being clones, they shared all their DNA with Bicone 39.16. However, due to some circumstances, they had slightly different fitness scores. Bicone 23 managed a very respectable 5.0117 fitness score and was ranked 2nd in the generation. But not to be outdone Bicone 22 managed to score a 5.17014 and ended up being the second-best performing individual of all time. Being so successful, the two bicones ended up producing 8 children between the two groups.
Bicone 23 was the first to crossover and had 2 children with its partner. These were Bicones 4 and 5. Bicone 4 was ranked 39th with a fitness score of 4.60251 and still shared the DNA of its second side with its grandparent Bicone 38.17 as well as most of its first side with Bicone 38.11 {2.32499, 79.663, -0.00213879, 0.192602} {0.308809, 39.9608, -0.0084247, 0.40629}. Bicone 5 on the other hand, was ranked 14th with a fitness score of 4.80535 and it still shared a lot of DNA with its grandparents {6.16084,53.0851,-0.00224948,0.0534469} {0.966617,39.6742,-0.0084247,0.40629}.
Bicone 22 had 6 children of its own with various other Bicones. These were; Bicone 8, ranked 11th with a fitness Score of 4.81784 and DNA {2.32499, 75.9855, -0.00224948, 0.192602} {0.966617, 39.6742, -0.00320023, 0.213833}; Bicone 9, ranked 7th with a fitness score of 4.88966 and DNA {6.10508, 79.663, -0.000594616, 0.0351901} {0.308809, 42.4246, -0.0084247, 0.40629}; Bicone 12, ranked 12th with a fitness score of 4.81705 and DNA {6.42695, 75.9855, 0.0015434, -0.107765} {0.308809, 39.6742, -0.00320023, 0.213833}; Bicone 13, ranked 2nd with a fitness score of 5.0344 and DNA {2.32499, 79.663, -0.00224948, 0.192602} {0.966617, 42.4246, -0.0084247, 0.40629}; Bicone 34 ranked 3rd with a fitness score of 4.99864 and DNA {0.66148, 73.5522, -0.000594616, 0.00582814} {0.966617, 39.6742, -0.0084247, 0.40629}; and finally Bicone 35, ranked 22nd with fitness score 4.74955 and DNA {2.32499, 79.663, -0.00224948, 0.192602} {0.308809, 42.4246, -0.0084247, 0.40629}
Bellow, I have attached the rainbow plot with the parameters occupied by individual 4 in gen 41 which was again ranked 39th in that generation. From this, we can see that while in its own generation it was a poor performer, overall it was upper middle of the pack. However, because of the density of other better performing antennas in this region, it is hard to distinguish which genes in this antenna are contributing the most to the drop in fitness score compared to its siblings and parents.
*Gene originating from Bicone 38.11
**Gene originating from Bicone 38.17
***Gene originating from Bicone 38.11 and 38.17 that is shared between the two. |
|
166
|
Tue Jun 21 13:20:41 2022 |
Ryan Debolt | Multigenerational Narrative draft | The story of individual 8 from generation 18. (draft)
Once, there was a curved bicone named individual 8 who was from generation 18. In many ways, it was similar to many other bicones but in fact, this individual was the best bicone that ever lived with a fitness score of 5.22097. It achieved this by having the following parameters; an inner radius of 6.31483, and 3.97024; lengths of 77.6017 and 40.9244; quadratic coefficients of, 0.00260615 and, -0.0103313; and finally, a linear coefficient of -0.197494 and
0.36119. Unfortunately, individual 8’s lineage ended with it as it never met that special someone (crossover) and didn't survive into the next generation (reproduction) despite having a 77.7% of doing at least one of these.
This bicone had a sibling with a fitness Score of 4.65861 and had the following parameters: 6.05614,79.663,-0.000353038,0.0331969 (side 1)
1.92878,37.9849,-0.00213265,0.156724 (side 2).
Unfortunately, this bicone also suffered the same fate as its sibling and failed to leave a lasting mark on this hypothetical world.
These two individuals had parents from generation 17 whose name’s were individuals 39 and individual 8. After having individuals 8 and 9 in generation 18, individuals 39 and 8 from generation had a nasty divorce (possibly leading to 8 and 9’s aversion to marriage) and remarried. In these marriages, they both produced two more children that would be our previous antenna’s step-siblings.
Antenna 8 from gen 17 married antenna 46 to produce antenna 20 with a fitness score of 4.57122, and antenna 21 with a fitness score of 4.71056. Individual 20 shared the same genes for the second set of linear and quadratic coefficients as its more successful step-sibling individual 8. 21 had no similarities to individual 8 but had 4 children in generation 19.
Antenna 39 from gen 17 married antenna 5 to produce antenna 28 with a fitness score of 4.71808 and antenna 29 with a fitness score of 4.90119. Antenna 28 shared had an identical side 1 to individual 8 and also shared the same inner radius and length on the second side, and had 2 children in generation 19. Individual 29 had no similarities to individual 8 but really got around and had 8 children with various partners in generation 19. |
|
120
|
Mon Nov 23 18:02:40 2020 |
Ryan Debolt | Monday Updates |
| Alex M |
Kept working with Amy, Alex P, Julie, and Ben on the AraSim fix. We fixed our issue from last week but have a new one in stage 2. It looks like the issue has to do with resetting the values for V_forfft right before stage 2 (around line 963). Check here for the current version of Report.cc: /users/PAS0654/pattonalexo/EFieldProject/11_23_20 |
| Ryan |
Fixed the issue with the segmentation faults in the GA (simple fix one line changed) and worked with Kai to start creating a spreadsheet of results. Results seem to suggest that 2 tournament to 8 roulette seems to be the ideal selection ratio and the initial test seem to suggest a small amount of reproduction is ideal but more testing is needed to confirm this. |
| |
|
| |
|
| |
|
| |
|
| |
|
|
|
122
|
Mon Nov 30 17:00:31 2020 |
Ethan Fahimi | Monday Updates |
| Alex M |
Kicked the loop back up. Helped Ethan and Parker with their projects during the working meeting. |
| Ryan |
Changed the tournament/roulette ratio, reproduction_no, and crossover_no to be read in variables in the GA to increase the quality of life when running the algorithm and to prep the code for some testing. Format for how to call the code is written at the top of the cpp program file. |
| Ethan F |
Worked with Alex M on further fixing AREA. The first generation works now, we are making minor fixes to get subsequent generations up and running. |
| |
|
| |
|
| |
|
| |
|
|
|
215
|
Mon May 1 05:45:02 2023 |
Alex M | Midscale PUEO Run | I've started a run of the PUEO loop to get some data for evolving the antenna width and height. We are running with 24 individuals, each with 400k neutrinos at an energy exponent of 19. Here is the output directory: /fs/ess/PAS1960/HornEvolutionOSC/GENETIS_PUEO/BiconeEvolution/current_antenna_evo_build/XF_Loop/Evolutionary_Loop/Run_Outputs/2023_05_01_Test_5
It's very important to note that the cross-pol data being used still comes from the data already stored in pueosim. This is because our extremely low cross polarizations were causing us to have no effective volumes. There's likely some issue in what values from XF we are choosing as our crosspols. |
| Attachment 1: run_details.txt
|
####### VARIABLES: LINES TO CHECK OVER WHEN STARTING A NEW RUN ###############################################################################################
RunName='2023_05_01_Test_5' ## This is the name of the run. You need to make a unique name each time you run.
TotalGens=10 ## number of generations (after initial) to run through
NPOP=24 ## number of individuals per generation; please keep this value below 99
Seeds=10 ## This is how many AraSim jobs will run for each individual## the number frequencies being iterated over in XF (Currectly only affects the output.xmacro loop)
FREQ=60 ## the number frequencies being iterated over in XF (Currectly only affects the output.xmacro loop)
NNT=40000 ## Number of Neutrinos Thrown in AraSim
exp=19 ## exponent of the energy for the neutrinos in AraSim
ScaleFactor=1.0 ## ScaleFactor used when punishing fitness scores of antennae larger than the drilling holes
GeoFactor=1 ## This is the number by which we are scaling DOWN our antennas. This is passed to many files
num_keys=6 ## how many XF keys we are letting this run use
database_flag=0 ## 0 if not using the database, 1 if using the database
## These next 3 define the symmetry of the cones.
PUEO=1 ## IF 1, we evolve the PUEO quad-ridged horn antenna, if 0, we evolve the Bicone
RADIUS=1 ## If 1, radius is asymmetric. If 0, radius is symmetric
LENGTH=1 ## If 1, length is asymmetric. If 0, length is symmetric
ANGLE=1 ## If 1, angle is asymmetric. If 0, angle is symmetric
CURVED=1 ## If 1, evolve curved sides. If 0, sides are straight
A=1 ## If 1, A is asymmetric
B=1 ## If 1, B is asymmetric
SEPARATION=0 ## If 1, separation evolves. If 0, separation is constant
NSECTIONS=2 ## The number of chromosomes
DEBUG_MODE=0 ## 1 for testing (ex: send specific seeds), 0 for real runs
## These next variables are the values passed to the GA
REPRODUCTION=6 ## Number (not fraction!) of individuals formed through reproduction
CROSSOVER=10 ## Number (not fraction!) of individuals formed through crossover
MUTATION=1 ## Probability of mutation (divided by 100)
SIGMA=5 ## Standard deviation for the mutation operation (divided by 100)
ROULETTE=2 ## Percent of individuals selected through roulette (divided by 10)
TOURNAMENT=2 ## Percent of individuals selected through tournament (divided by 10)
RANK=6 ## Percent of individuals selected through rank (divided by 10)
ELITE=0 ## Elite function on/off (1/0)
|
|
161
|
Tue Jun 7 14:14:21 2022 |
Dylan Wells | Matching Circuits Slides | Slides contatining my notes on matching circuits.
https://docs.google.com/presentation/d/1x25nhiqaW7LvPZ1pNZ5O4ZzsWZbtgqxBQ5haB9uWgQY/edit?usp=sharing |
|
Draft
|
Sat Apr 4 11:04:28 2026 |
Dylan Wells | Matching Circuits Slides | Slides contatining my notes on matching circuits.
https://docs.google.com/presentation/d/1x25nhiqaW7LvPZ1pNZ5O4ZzsWZbtgqxBQ5haB9uWgQY/edit?usp=sharing |
|
191
|
Mon Feb 6 10:23:53 2023 |
Dylan Wells | Matching Circuit Parts | Attached is a spreadsheet with the information on the parts we need for the N=14 matching circuit board.
https://docs.google.com/spreadsheets/d/1x8dX3tNE-WSHjH_slj_EH4XsHcAnCVC05XiRBFPtIUc/edit#gid=0 |
|
137
|
Wed Sep 8 16:36:33 2021 |
Alex M | MODE Workshop Presentation | We presented at a workshop put on by the MODE collaboration. MODE is a collaboration dedicated to applying Automatic Differentiation to detector design. Here's the website: https://mode-collaboration.github.io/#:~:text=MODE%20(for%20Machine%2Dlearning%20Optimized,in%20space%2C%20and%20in%20nuclear
|
| Attachment 1: GENETIS_MODE_Presentation.pptx
|
|
142
|
Fri Feb 4 16:50:09 2022 |
Ryan Debolt | Loop Run |
- Run Type
- Run Date
- Run Name
- Why are we doing this run?
- To test rank selection in main loop
- What is different about this run from the last?
- Rank Slection is being used.
- Parents.csv introduced.
- Elite is being turned off.
- Symmetric, asymmetric, linear, nonlinear (what order):
- Number of individuals (NPOP):
- Number of neutrinos thrown in AraSim (NNT):
- Operatiors used (% of each):
- 06% Reproduction
- 72% Crossver
- 22% Immigration
- 1% M_rate (unused)
- 5% sigma (unused)
- Selection methods used (% of each):
- 0% Elite
- 0% Reproduction
- 10% Tournament
- 90% Rank
- Are we using the database?
Directory: /fs/ess/PAS1960/BiconeEvolutionOSC/BiconeEvolution/current_antenna_evo_build/XF_Loop/Evolutionary_Loop/Run_Outputs/2022_02_04_Rank |
| Attachment 1: run_details.txt
|
####### VARIABLES: LINES TO CHECK OVER WHEN STARTING A NEW RUN ###############################################################################################
RunName='2022_02_04_Rank' ## This is the name of the run. You need to make a unique name each time you run.
TotalGens=100 ## number of generations (after initial) to run through
NPOP=50 ## number of individuals per generation; please keep this value below 99
Seeds=10 ## This is how many AraSim jobs will run for each individual## the number frequencies being iterated over in XF (Currectly only affects the output.xmacro loop)
FREQ=60 ## the number frequencies being iterated over in XF (Currectly only affects the output.xmacro loop)
NNT=30000 ## Number of Neutrinos Thrown in AraSim
exp=18 ## exponent of the energy for the neutrinos in AraSim
ScaleFactor=1.0 ## ScaleFactor used when punishing fitness scores of antennae larger than the drilling holes
GeoFactor=1 ## This is the number by which we are scaling DOWN our antennas. This is passed to many files
num_keys=4 ## how many XF keys we are letting this run use
database_flag=0 ## 0 if not using the database, 1 if using the database
## These next 3 define the symmetry of the cones.
RADIUS=1 ## If 1, radius is asymmetric. If 0, radius is symmetric
LENGTH=1 ## If 1, length is asymmetric. If 0, length is symmetric
ANGLE=1 ## If 1, angle is asymmetric. If 0, angle is symmetric
CURVED=1 ## If 1, evolve curved sides. If 0, sides are straight
A=1 ## If 1, A is asymmetric
B=1 ## If 1, B is asymmetric
SEPARATION=0 ## If 1, separation evolves. If 0, separation is constant
NSECTIONS=2 ## The number of chromosomes
DEBUG_MODE=0 ## 1 for testing (ex: send specific seeds), 0 for real runs
## These next variables are the values passed to the GA
REPRODUCTION=3 ## Number (not fraction!) of individuals formed through reproduction
CROSSOVER=36 ## Number (not fraction!) of individuals formed through crossover
MUTATION=1 ## Probability of mutation (divided by 100)
SIGMA=5 ## Standard deviation for the mutation operation (divided by 100)
ROULETTE=0 ## Percent of individuals selected through roulette (divided by 10)
TOURNAMENT=1 ## Percent of individuals selected through tournament (divided by 10)
RANK=9 ## Percent of individuals selected through rank (divided by 10)
ELITE=0 ## Elite function on/off (1/0)
#####################################################################################################################################################
|
|
128
|
Fri Jan 29 15:59:21 2021 |
Alex M | Loop Demonstration Video | Here's the video I made a few months ago demonstrating how to run the loop. There might be some changes that have been made since then, but they'll generally be minor so it should still be representative of what running the loop looks like. It might need to be shared with you through the Microsoft drive service OSU gives us in order to view it, so either message me on Slack or email me at machtay.1@osu.edu and I'll share the link through there.
https://drive.google.com/file/d/1yL_eH_w2hXNt7J7YOwS5xl3ZHwGyl5JR/view |
|
5
|
Thu Feb 21 14:07:14 2019 |
Julie Rolla | Logging into Nutau via XRDP | As mentioned in the manual, using ssh to login to Nutau causes delays; the XFdtd GUI cannot be surpress and must be forwarded via X11 forwarding, and this is extremely slow. We are currently looking into using a newly suggested option, XRDP. Information will be added as this process continues. |
|
236
|
Wed Aug 2 12:49:57 2023 |
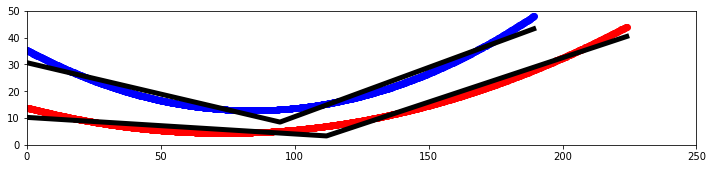
Dylan Wells | Line of Best-Fit Straightened Sides Antenna | Previously, we have tried to straighten the sides of the best-evolved curved antenna in elog 229. However, there were potential issues with how closely this line resembled the curve of the antenna.
So, I attempted to create another straightened sides antenna using a linear regression model to find the best fitting line for the curve to create an antenna.
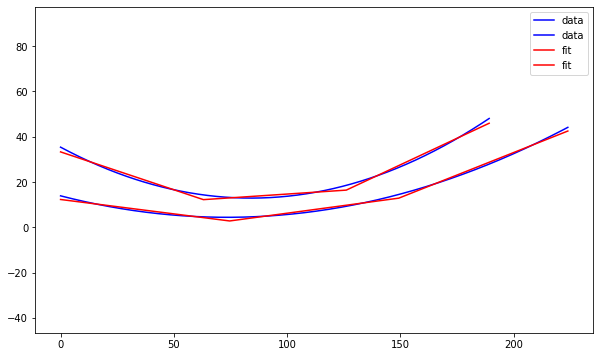
I made a Python notebook to separate the equations for each curve into 1000 discrete points. Then I ran a linear regression model to fit a curve of the points 1 - n, and a second curve of the n - 1000 points, looping from n = 2 to n = 999.
These results are from the combined output with the least squared error compared to the original.
Pictured is a plot showing the two sides of the curved bicone in red and blue, with the best fitting lines for each in black as well as a model in XF.
Results:
The antenna has a fitness score of 3.80627 with an error of 0.0759725.
This is much lower than the 5.71 of the curved antenna and 5.11 of the other attempt at straightening.
For the next attempt, we could consider constraining the endpoints to be the same as the original antenna to conserve the radius, and/or adding an extra line to fit the curve.
I've also attached a picture of what my notebook found for fitting 3 lines to the curve (not modeled or tested).
Professor Chen recommended using 3 sides and constraining the outer radii of the cones to match the original curved design.
Path to linear regression notebook: /users/PAS1960/dylanwells1629/developing/notebook.ipynb
Path to XF project: /users/PAS1960/dylanwells1629/straightened.xf |
| Attachment 1: output.png
|  |
| Attachment 2: image_(3).png
| .png) |
| Attachment 3: output.png
|  |
|
126
|
Tue Jan 19 19:00:30 2021 |
Alex M | Latest Run Details/Running on Slurm | OSC managed to figure out our issue with running XF from an interactive job on Slurm. Previously, we were losing our connection to x11 forwarding. The solution is to use sinteractive instead of srun to obtain an interactive job. Here's the syntax:
sinteractive -A PAS0654 -t <time> -N 1 -n 8 -p serial
The -p serial flag denotes the type of partition to request. It's important to specify this, as otherwise it will default to the debug node, which has a limit of 1 hour.
We're going to begin a new run. The title is Machtay_2021_1_15_NPOP50_Asym . It is using the latest version of the GA Ryan has been working on (Latest_GA_Asym.cpp). We're using 6% reproduciton, 72% crossover, and 22% mutation. We are using 80% roulette selection. This correlates to the three numbers passed into the GA being 3 36 8 (since we're using 50 individuals). |
|