| ID |
Date |
Author |
Subject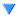 |
|
179
|
Tue Aug 16 11:41:07 2022 |
Dylan Wells | Instructions for Running IceMC | Running IceMC:
Go into the directory ../anitaBuildTool/build/components/icemc/
And run the command
*May need to chmod -R 775 ../anitaBuildTool/comonents/icemc/ if you get a permissions error
inputFile:
Must be the full path to the file
Config files are found in ../anitaBuildTool/components/icemc
Ex: /users/PAS1960/dylanwells1629/anitaBuildTool/components/icemc/inputs.anita4.conf
Config files are found in ../anitaBuildTool/components/icemc
outputDirectory:
Will be made in ../anitaBuildTool/build/components/icemc/ by default, specify full path otherwise.
runNumber:
The run number.
numberOfNeutrinos:
The number of neutrinos generated in the simulation.
Can be found in inputs.conf
Default is 2,000,000.
#How many neutrinos to generate
triggerThreshold:
Threshold for each band for the trigger.
Default is 2.3
#thresholds for each band- this is only for the frequency domain voltage trigger. If using a different trigger scheme then keep these at the default values of 2.3 because the max among them is used for the chance in hell cuts
energyExponent:
The exponent of the energy for the neutrinos
Can be found in input.conf
Default is 1020
# Select energy (just enter the exponent) or (30) for baseline ES&S (1) for E^-1 (2) for E^-2 (3) for E^-3 (4) for E^-4 (5) for ES&S flux with cosmological constant (6) for neutrino GZK flux from Iron nuclei (16-22)not using spectrum but just for a single energy (101-114)use all kinds of theoretical flux models |
|
Draft
|
Mon Apr 6 03:03:14 2026 |
Dylan Wells | Instructions for Running IceMC | Running IceMC:
Go into the directory ../anitaBuildTool/build/components/icemc/
And run the command
*May need to chmod -R 775 ../anitaBuildTool/comonents/icemc/ if you get a permissions error
inputFile:
Must be the full path to the file
Config files are found in ../anitaBuildTool/components/icemc
Ex: /users/PAS1960/dylanwells1629/anitaBuildTool/components/icemc/inputs.anita4.conf
Config files are found in ../anitaBuildTool/components/icemc
outputDirectory:
Will be made in ../anitaBuildTool/build/components/icemc/ by default, specify full path otherwise.
runNumber:
The run number.
numberOfNeutrinos:
The number of neutrinos generated in the simulation.
Can be found in inputs.conf
Default is 2,000,000.
#How many neutrinos to generate
triggerThreshold:
Threshold for each band for the trigger.
Default is 2.3
#thresholds for each band- this is only for the frequency domain voltage trigger. If using a different trigger scheme then keep these at the default values of 2.3 because the max among them is used for the chance in hell cuts
energyExponent:
The exponent of the energy for the neutrinos
Can be found in input.conf
Default is 1020
# Select energy (just enter the exponent) or (30) for baseline ES&S (1) for E^-1 (2) for E^-2 (3) for E^-3 (4) for E^-4 (5) for ES&S flux with cosmological constant (6) for neutrino GZK flux from Iron nuclei (16-22)not using spectrum but just for a single energy (101-114)use all kinds of theoretical flux models |
|
178
|
Wed Aug 10 22:38:20 2022 |
Dylan Wells | Instructions for Installing IceMC | Installing IceMC:
I: Getting anitaBuildTool
Clone the anitaBuildTool repository (https://github.com/anitaNeutrino/anitaBuildTool) to your user.
II: Get the Anita.sh file onto your user.
Either copy the file from /users/PAS1960/dylanwells1629/Anita.sh
or create a new file Anita.sh with the following code
# .bashrc
# Source global definitions
if [ -f /etc/bashrc ]; then
. /etc/bashrc
fi
#this modules were originally loading in both the env.sh, and bashrc_anita.sh files. This was redundant so it was added here, and removed from the others.
module load gnu/7.3.0
module load gnu
module load mvapich2
module load fftw3
#module load python/3.6-conda5.2
module load cmake
PATH=$PATH:$HOME/.local/bin:$home/bin
export PATH
export CC=`which gcc`
export CXX=`which g++`
export FFTWDIR=/fs/project/PAS0654/shared_software/fftw3/gnu/6.3/mvapich2/2.2/3.3.5
export ANITA_SOURCE_DIR=~/anitaBuildTool/
export ANITA_UTIL_INSTALL_DIR=~/anitaBuildTool/
export ICEMC_SRC_DIR=~/anitaBuildTool/components/icemc/
export ICEMC_BUILD_DIR=~/anitaBuildTool/build/components/icemc/
export DYLD_LIBRARY_PATH=${ICEMC_SRC_DIR}:${ICEMC_BUILD_DIR}:${DYLD_LIBRARY_PATH}
export ROOTSYS=/fs/project/PAS0654/shared_software/anita/owens_pitzer/build/root
# User specific aliases and functions
#This env.sh is for running the BiconeEvolution GENETIS software. This should only be un-commented if you are running GENETIS software. When you do this, comment out env.sh.
#source ~/new_root_setup.sh
source /fs/project/PAS0654/shared_software/anita/owens_pitzer/build/root/bin/thisroot.sh
#source /cvmfs/ara.opensciencegrid.org/trunk/centos7/setup.sh
#module load python/3.6-conda5.2
#BiconeGENETIS directory shortcut SHARED
alias GE='cd ../../../fs/project/PAS0654/BiconeEvolutionOSC/BiconeEvolution/current_antenna_evo_build/XF_Loop/Evolutionary_Loop/'
#emacs Alias
alias emacs='emacs -nw'
#root alias
alias root='root -l'
#Alias
alias l="ls"
alias python='/cvmfs/ara.opensciencegrid.org/trunk/centos7/misc_build/bin/python3.9'
Then source the file
Note: You will need access to PAS0654 for this step or you will get a permissions error.
III: Running the build tool.
Go into the anitaBuildTool directory
And run the building script
Note: There will be an error if you source files for running Ara in your .bashrc
Comment these out and restart your terminal before running the build. (remember to source Anita.sh before running the build tool. You could also source Anita.sh in your .bashrc)

Error if you source files for running Ara:
CMake Error at components/libRootFftwWrapper/cmake_install.cmake:238 (file):
file INSTALL cannot copy file
"/users/PAS1960/dylanwells1629/anitaBuildTool/components/libRootFftwWrapper/include/AnalyticSignal.h"
to
"/cvmfs/ara.opensciencegrid.org/v2.0.0/centos7/ara_build/include/AnalyticSignal.h":
Read-only file system.
Call Stack (most recent call first):
|
|
204
|
Mon Mar 20 16:31:09 2023 |
Alex M | Installing PUEOsim | Will sent me some info on how to install PUEOsim. I attached a markdown file he sent for how to install PUEOsim. He also sent the path to pre-compiled installation of PUEOsim. Here's the path to the pre-compiled installation: /users/PAS0654/wluszczak/PUEO_shared
Here is his full message, which details that you must also run set_env.sh before running PUEOsim:
a pre-compiled version is located at /users/PAS0654/wluszczak/PUEO_shared . Inside that directory, there's a set_env.sh script that you will need to source before running anything (hopefully this works, though there's a high probability you might run into permissions issues with the dependencies).
To run PUEOsim, use the following (see here for instructions: https://github.com/PUEOCollaboration/pueoSim):
./simulatePueo -i {inputFile} -o {outputDirectory} -r {runNumber} -n {numberOfNeutrinos} -e {energyExponent}
The outputs will be root files, but it should be possible (either modifying our own version directly or with a script) to print out the effective volume to a .txt to read.
|
| Attachment 1: CentOS_7_install.md
|
#pueoBuilder Installation Tutorial
This tutorial will guide you through the process of building the tools included in pueoBuilder from scratch, including the prerequisites and any environment variables that you will need to set. This sort of thing is always a bit of a nightmare process for me, so hopefully this guide can help you skip some of the frustration that I ran into. I did not have root acces on the system I was building on, so the instructions below are what I had to do to get things working with local installations. If you have root access, then things might be a bit easier. For reference I'm working on CentOS 7, other operating systems might have different problems that arise.
##Prerequisites
As far as I can tell, the prerequisites that need to be built first are:
-cmake 3.21.2 (I had problems with 3.11.4)
-gcc 11.1.0 (9.X will not work)
-fftw 3.3.9
-gsl 2.7.1 (for ROOT)
-ROOT 6.24.00
###CMake
You can download the source files for CMake here: https://cmake.org/download/. Untar the source files with:
tar -xzvf cmake-3.22.1.tar.gz
Compiling CMake is as easy as following the directions on the website: https://cmake.org/install/, but since we're doing a local build, we'll use the `configure` script instead of the listed `bootstrap` script. As an example, suppose that I downloaded the above tar file to `/scratch/wluszczak/cmake`:
mkdir install
cd cmake-3.22.1
./configure --prefix=/scratch/wluszczak/cmake/install
make
make install
You should additionally add this directory to your `$PATH` variable:
export PATH=/scratch/wluszczak/cmake/install/bin:$PATH
To check to make sure that you are using the correct version of CMake, run:
cmake --version
and you should get:
cmake version 3.22.1
CMake suite maintained and supported by Kitware (kitware.com/cmake).
### gcc 11.1.0
Download the gcc source from github here: https://github.com/gcc-mirror/gcc/tags. I used the 11.1.0 release, though there is a more recent 11.2.0 release that I have not tried. Once you have downloaded the source files, untar the directory:
tar -xzvf gcc-releases-gcc-11.1.0.tar.gz
Then install the prerequisites for gcc:
cd gcc-releases-gcc-11.1.0
contrib/download_prerequisites
One of the guides I looked at also recommended installing flex separately, but I didn't seem to need to do this, and I'm not sure how you would go about it without root priviledges, though I imagine it's similar to the process for all the other packages here (download the source and then build by providing an installation prefix somewhere)
After you have installed the prerequisites, create a build directory:
cd ../
mkdir build
cd build
Then configure GCC for compilation like so:
../gcc-releases-gcc-11.1.0/configure -v --prefix=/home/wluszczak/gcc-11.1.0 --enable-checking=release --enable-languages=c,c++,fortran --disable-multilib --program-suffix=-11.1
I don't remember why I installed to my home directory instead of the /scratch/ directories used above. In principle the installation prefix can go wherever you have write access. Once things have configured, compile gcc with:
make -j $(nproc)
make install
Where `$(nproc)` is the number of processing threads you want to devote to compilation. More threads will run faster, but be more taxing on your computer. For reference, I used 8 threads and it took ~15 min to finish.
Once gcc is built, we need to set a few environment variables:
export PATH=/home/wluszczak/gcc-11.1.0/bin:$PATH
export LD_LIBRARY_PATH=/home/wluszczak/gcc-11.1.0/lib64:$LD_LIBRARY_PATH
We also need to make sure cmake uses this compiler:
export CC=/home/wluszczak/gcc-11.1.0/bin/gcc-11.1
export CXX=/home/wluszczak/gcc-11.1.0/bin/g++-11.1
export FC=/home/wluszczak/gcc-11.1.0/bin/gfortran-11.1
If your installation prefix in the configure command above was different, substitute that directory in place of `/home/wluszczak/gcc-11.1.0` for all the above export commands. To easily set these variables whenever you want to use gcc-11.1.0, you can stick these commands into a single shell script:
#load_gcc11.1.sh
export PATH=/home/wluszczak/gcc-11.1.0/bin:$PATH
export LD_LIBRARY_PATH=/home/wluszczak/gcc-11.1.0/lib64:$LD_LIBRARY_PATH
export CC=/home/wluszczak/gcc-11.1.0/bin/gcc-11.1
export CXX=/home/wluszczak/gcc-11.1.0/bin/g++-11.1
export FC=/home/wluszczak/gcc-11.1.0/gfortran-11.1
(again substituting your installation prefix in place of mine). You can then set all these environment variables by simply running:
source load_gcc11.1.sh
Once this is done, you can check that gcc-11.1.0 is properly installed by running:
gcc-11.1 --version
Note that plain old
gcc --version
might still point to an older version of gcc. This is fine though.
###FFTW 3.3.9
Grab the source code for the appropriate versino of FFTW from here: http://www.fftw.org/download.html
However, do NOT follow the installation instructions on the webpage. Those instructions might work if you have root privileges, but I personally couldn't seem to to get things to work that way. Instead, we're going to build fftw with cmake. Untar the fftw source files:
tar -xzvf fftw-3.3.9.tar.gz
Make a build directory and cd into it:
mkdir build
cd build
Now build using cmake, using the flags shown below. For reference, I downloaded and untarred the source file in `/scratch/wluszczak/fftw/build`, so adjust your install prefix accordingly to point to your own build directory that you created in the previous step.
cmake -DCMAKE_INSTALL_PREFIX=/scratch/wluszczak/fftw/build/ -DBUILD_SHARED_LIBS=ON -DENABLE_OPENMP=ON -DENABLE_THREADS=ON ../fftw-3.3.9
make install -j $(nproc)
Now comes the weird part. Remove everything except the `include` and `lib64` directories in your build directory (if you installed to a different `CMAKE_INSTALL_PREFIX`, the include and lib64 directories might be located there instead. The important thing is that you want to remove everything, but leave the `include` and `lib64` directories untouched):
rm *
rm -r CMakeFiles
Now rebuild fftw, but with an additional flag:
cmake -DCMAKE_INSTALL_PREFIX=/scratch/wluszczak/fftw/build/ -DBUILD_SHARED_LIBS=ON -DENABLE_OPENMP=ON -DENABLE_THREADS=ON -DENABLE_FLOAT=ON ../fftw-3.3.9
make install -j $(nproc)
At the end of the day, your fftw install directory should have the following files:
include/fftw3.f
include/fftw3.f03
include/fftw3.h
include/fftw3l.f03
include/fftw3q.f03
lib64/libfftw3f.so
lib64/libfftw3f_threads.so.3
lib64/libfftw3_omp.so.3.6.9
lib64/libfftw3_threads.so
lib64/libfftw3f_omp.so
lib64/libfftw3f.so.3
lib64/libfftw3f_threads.so.3.6.9
lib64/libfftw3.so
lib64/libfftw3_threads.so.3
lib64/libfftw3f_omp.so.3
lib64/libfftw3f.so.3.6.9
lib64/libfftw3_omp.so
lib64/libfftw3.so.3
lib64/libfftw3_threads.so.3.6.9
lib64/libfftw3f_omp.so.3.6.9
lib64/libfftw3f_threads.so
lib64/libfftw3_omp.so.3
lib64/libfftw3.so.3.6.9
Why do we have to do things this way? I don't know, I'm bad at computers. Maybe someone more knowledgeable knows. I found that when I didn't do this step, I'd run into errors that pueoBuilder could not find some subset of the required files (either the ones added by building with `-DENABLE_FLOAT`, or the ones added by building without `-DENABLE_FLOAT`).
Once fftw has been installed, export your install directory (the one with the include and lib64 folders) to the following environment variable:
export FFTWDIR=/scratch/wluszczak/fftw/build
Again, substituting your own fftw install prefix that you used above in place of `/scratch/wluszczak/fftw/build`
###gsl 2.7.1
gsl 2.7.1 is needed for the `mathmore` option in ROOT. If you have an outdated version of gsl, ROOT will still compile, but it will skip installing `mathmore` and `root-config --has-mathmore` will return `no`. To fix this, grab the latest source code for gsl from here: https://www.gnu.org/software/gsl/. Untar the files to a directory of your choosing:
tar -xzvf gsl-latest.tar.gz
For some reason I also installed gsl to my home directory, but in principle you can put it wherever you want.
mkdir /home/wluszczak/gsl
./configure --prefix=/home/wluszczak/gsl
make
make check
make install
To make sure ROOT can find this installation of gsl, you'll again need to set an environment variable prior to building ROOT:
export GSL_ROOT_DIR=/home/wluszczak/gsl/
I also added this to my $PATH variable, though I don't remember if that was required to get things working or not:
export PATH=/home/wluszczak/gsl/bin/:$PATH
export LD_LIBRARY_PATH=/home/wluszczak/gsl/lib:$LD_LIBRARY_PATH
###ROOT 6.24.00
Download the specific version of ROOT that you need from here: https://root.cern/install/all_releases/
You might need to additionally install some of the dependencies (https://root.cern/install/dependencies/), but it seems like everything I needed was already installed on my system.
Untar the source you downloaded:
tar -xzvf root_v6.24.00.source.tar.gz
Make some build and install directories:
mkdir build install
cd build
Run CMake, but be sure to enable the fortan, mathmore and minuit2 options. For reference, I had downloaded and untarred the source files to `/scratch/wluszczak/root`. Your installation and source paths will be different.
cmake -DCMAKE_INSTALL_PREFIX=/scratch/wluszczak/root/install/ /scratch/wluszczak/root/root-6.24.00/ -Dfortran=ON -Dminuit2=ON -Dmathmore=ON
Note: if you end up with an error related to compiling XROOTD, then add -Dxrootd=OFF to the original cmake command above.
Then proceed to start the build:
cmake --build . --target install -j $(nproc)
If everything has worked then after the above command finishes running, you should be able to run the following file to finish setting up ROOT:
source ../install/bin/thisroot.sh
##pueoBuilder
By this point, you should have working installations of CMake 3.21.2, gcc-11.1.0, fftw 3.3.9, and ROOT 6.24.00. Additionally, the following environment variables should have been set:
export PATH=/scratch/wluszczak/cmake/install/bin:$PATH
export PATH=/home/wluszczak/gcc-11.1.0/bin:$PATH
export LD_LIBRARY_PATH=/home/wluszczak/gcc-11.1.0/lib64:$LD_LIBRARY_PATH
export CC=/home/wluszczak/gcc-11.1.0/bin/gcc-11.1
export CXX=/home/wluszczak/gcc-11.1.0/bin/g++-11.1
export FC=/home/wluszczak/gcc-11.1.0/gfortran-11.1
export FFTWDIR=/scratch/wluszczak/fftw/build
At this point, the hard work is mostly done. Check out pueoBuilder with:
git clone git@github.com:PUEOCollaboration/pueoBuilder
set the following environment variables:
export PUEO_BUILD_DIR=/scratch/wluszczak/PUEO/pueoBuilder
export PUEO_UTIL_INSTALL_DIR=/scratch/wluszczak/PUEO/pueoBuilder
export NICEMC_SRC=${PUEO_BUILD_DIR}/components/nicemc
export NICEMC_BUILD=${PUEO_BUILD_DIR}/build/components/nicemc
export PUEOSIM_SRC=${PUEO_BUILD_DIR}/components/pueoSim
export LD_LIBRARY_PATH=${PUEO_UTIL_INSTALL_DIR}/lib:$LD_LIBRARY_PATH
Where $PUEO_BUILD_DIR and $PUEO_UTIL_INSTALL_DIR point to where you cloned pueoBuilder to (in my case, `/scratch/wluszczak/PUEO/pueoBuilder`. Now you should be able to just run:
./pueoBuilder.sh
Perform a prayer to the C++ gods while you're waiting for it to compile, and hopefully at the end of the day you'll have a working set of PUEO software.
##Issues I Ran Into
If you already have an existing installation of ROOT, you may still need to recompile to make sure you're using the same c++ standard that the PUEO software is using. I believe the pre-compiled ROOT binaries available through their website are insufficient, though maybe someone else has been able to get those working.
If you're running into errors about c++ standard or compiler version even after you have installed gcc-11.1.0, then for some reason your system isn't recognizing your local installation of gcc-11.1.0. Check the path variables ($PATH and $LD_LIBRARY_PATH) to make sure the gcc-11.1.0 `bin` directory is being searched.
If you're running into an error that looks like:
CMake Error: The following variables are used in this project, but they are set to NOTFOUND.
Please set them or make sure they are set and tested correctly in the CMake files:
FFTWF_LIB (ADVANCED)
then pueoBuilder can't seem to find your fftw installation (or files that are supposed to be included in that installation), try rebuilding with different flags according to which files it seems to think are missing.
If it seems like pueoBuilder can't seem to find your fftw installation at all (i.e. you're getting some error that looks like `missing: FFTW_LIBRARIES` or `missing: FFTW_INCLUDES`), check the environment variables that are supposed to point to your fftw installation (`$FFTWDIR`) and make sure there are the correct files in the `lib` and `include` subdirectories.
|
|
157
|
Mon Jun 6 14:19:59 2022 |
Alex M | Important Runs (2) | This is a duplicate post of a previous post from the end of 2020 where I listed the important runs with some of their details in a table (as below). I am extending this table to include the important runs that have been conducted since this post. This includes the run used for the paper as well as the curved run done earlier this year.
In order to access the data for these runs, you can find them by going to this directory: /fs/ess/PAS1960/BiconeEvolutionOSC/BiconeEvolution/current_antenna_evo_build/XF_Loop/Evolutionary_Loop/Run_Outputs
Some of these runs can also be accessed in the old project space directory, though they should all be contained in the above directory. Here's the path if interested:
The runs are contained in directories available in the above path. Use caution when listing files in some of these directories--some contain many files (primarily the .uan files -- more recent runs are better organized), which means it may take a long time to list files/directories.
| Name |
Description |
Symmetry |
NPOP |
Generations |
Roulette/Tourney/Rank |
Crossover |
Reproduction |
Mutation |
Injection |
Penalty |
Neutrinos |
|
| Machtay_20200824_Real_Run |
First real run with significant amounts of data after the summer improvements.______________________________________
|
Symmetric |
10 |
15 |
100% Roulette |
100% |
0% |
- |
100% |
Yes |
100k |
|
| Machtay_20200827_Asym_Length_Run |
First asymmetric length run after summer improvements. |
Asymmetric length |
10 |
17 |
100% Roulette |
100% |
0% |
- |
100% |
Yes |
100k |
|
| Machtay_20200831_Asym_Length_and_Angle |
Asymmetric length and angle run after summer improvements. |
Asymmetric length and angle |
10 |
42 |
100% Roulette |
100% |
0% |
- |
100% |
Yes |
150k |
|
| Machtay_20200911_Symmetric |
Longer symmetric run with fewer neutrinos. |
Symmetric |
10 |
35 |
100% Roulette |
100% |
0% |
- |
100% |
Yes |
30k |
|
| Machtay_20200914_Asymmetric_50_Individuals |
Longer asymmetric run with fewer neutrinos. |
Asymmetric (all dimensions) |
50 |
26 |
100% Roulette |
100% |
0% |
- |
100% |
Yes |
30k |
|
| Machtay_20201016_Symmetric_Improved_GA |
First run using improvements to GA based on Ryan's paperclip/fast loop analysis. |
Symmetric |
50 |
10 |
50/50/0 |
75% |
10% |
- |
15% |
Yes |
30k |
|
| Machtay_20201023_300K_Nus_50_Individuals |
Started with all identical individuals to demonstrate evolution; replaced penalty with hard cutoff. Increased Nus for higher fitness score precision. |
Asymmetric (all dimensions |
50 |
25 |
50/50/0 |
75% |
10% |
- |
15% |
No |
300k |
|
| AraSim_Polarity_Fix_2021_03_19 |
Run used in the paper. In this run, we fixed an error that had been noticed by Brian and Jorge in AraSim. The error involved the polarity of the signals in Report.cc (hence the name of this run). |
Asymmetric(all dimensions) |
50 |
31 |
80/20/0 |
72% |
6% |
- |
22% |
No |
300k |
|
| 2022_02_08_Rank_Test |
This was the first long run done using a new gene for the curvature of the cones. We recast the side lengths to be described by the coefficients of a quadratic polynomial, rather than by the opening angle. This also used rank selection instead of roulette.
Additionally, mutation has been changed here so as to apply small perturbations to existing genes rather than regenerating those genes altogether. This only applies to individuals created by crossover. The mutation column indicates the probability of mutating a gene and the standard deviation of the gaussian that determines the change (in terms of % of the original value).
|
Asymmetric, Quadratic |
50 |
50 |
0/90/10 |
72% |
6% |
1%, 5% |
22% |
No |
300k |
|
| 2022_04_14_Identical_Asym_Lower_Min |
This run used the asymmetric GA to see if by lowering the minimum length (down to 10 cm instead of 37.5) the GA would try to run away to ever smaller lengths. |
Asymmetric |
50 |
6 |
2/8/0 |
72% |
6% |
- |
22% |
No |
300k |
|
|
|
124
|
Mon Dec 7 22:51:08 2020 |
Alex M | Important Runs | Today I removed some of the run directories which had very little or no data or weren't worth keeping around. There are still a few that I think can be removed, but I'm keeping them until we can get a consensus that they can definitely be removed. Below I listed the names and descriptions of the runs that I think we should definitely preserve going forward. In general, the more data contained in the run directory, the more important it is to keep around.
| Name |
Description |
Symmetry |
NPOP |
Generations |
Roulette/Tourney |
Crossover |
Reproduction |
Mutation |
Penalty |
Neutrinos |
|
| Machtay_20200824_Real_Run |
First real run with significant amounts of data after the summer improvements.______________________________________
|
Symmetric |
10 |
15 |
100% Roulette |
100% |
0% |
100% |
Yes |
100k |
|
| Machtay_20200827_Asym_Length_Run |
First asymmetric length run after summer improvements. |
Asymmetric length |
10 |
17 |
100% Roulette |
100% |
0% |
100% |
Yes |
100k |
|
| Machtay_20200831_Asym_Length_and_Angle |
Asymmetric length and angle run after summer improvements. |
Asymmetric length and angle |
10 |
42 |
100% Roulette |
100% |
0% |
100% |
Yes |
150k |
|
| Machtay_20200911_Symmetric |
Longer symmetric run with fewer neutrinos. |
Symmetric |
10 |
35 |
100% Roulette |
100% |
0% |
100% |
Yes |
30k |
|
| Machtay_20200914_Asymmetric_50_Individuals |
Longer asymmetric run with fewer neutrinos. |
Asymmetric (all dimensions) |
50 |
26 |
100% Roulette |
100% |
0% |
100% |
Yes |
30k |
|
| Machtay_20201016_Symmetric_Improved_GA |
First run using improvements to GA based on Ryan's paperclip/fast loop analysis. |
Symmetric |
50 |
10 |
50/50 |
75% |
10% |
15% |
Yes |
30k |
|
| Machtay_20201023_300K_Nus_50_Individuals |
Started with all identical individuals to demonstrate evolution; replaced penalty with hard cutoff. Increased Nus for higher fitness score precision. |
Asymmetric (all dimensions |
50 |
25 |
50/50 |
75% |
10% |
15% |
No |
300k |
|
| |
|
|
|
|
|
|
|
|
|
|
|
| |
|
|
|
|
|
|
|
|
|
|
|
|
|
152
|
Fri Apr 8 16:07:22 2022 |
Alex M | Identical Asymmetric Lowered Length Minimum Run | We are trying to do a short run of the asymmetric bicone so that we can see how it tries to evolve the antenna when the minimum length is lowered from 37.5 cm to 10 cm. Currently, there is a problem with the loop in the asymmetric version. Attached is the run detail file. |
| Attachment 1: run_details.txt
|
####### VARIABLES: LINES TO CHECK OVER WHEN STARTING A NEW RUN ###############################################################################################
RunName='2022_04_07_Identical_Asym' ## This is the name of the run. You need to make a unique name each time you run.
TotalGens=100 ## number of generations (after initial) to run through
NPOP=50 ## number of individuals per generation; please keep this value below 99
Seeds=10 ## This is how many AraSim jobs will run for each individual## the number frequencies being iterated over in XF (Currectly only affects the output.xmacro loop)
FREQ=60 ## the number frequencies being iterated over in XF (Currectly only affects the output.xmacro loop)
NNT=30000 ## Number of Neutrinos Thrown in AraSim
exp=18 ## exponent of the energy for the neutrinos in AraSim
ScaleFactor=1.0 ## ScaleFactor used when punishing fitness scores of antennae larger than the drilling holes
GeoFactor=1 ## This is the number by which we are scaling DOWN our antennas. This is passed to many files
num_keys=4 ## how many XF keys we are letting this run use
database_flag=0 ## 0 if not using the database, 1 if using the database
## These next 3 define the symmetry of the cones.
RADIUS=1 ## If 1, radius is asymmetric. If 0, radius is symmetric
LENGTH=1 ## If 1, length is asymmetric. If 0, length is symmetric
ANGLE=1 ## If 1, angle is asymmetric. If 0, angle is symmetric
CURVED=0 ## If 1, evolve curved sides. If 0, sides are straight
A=1 ## If 1, A is asymmetric
B=1 ## If 1, B is asymmetric
SEPARATION=0 ## If 1, separation evolves. If 0, separation is constant
NSECTIONS=2 ## The number of chromosomes
DEBUG_MODE=0 ## 1 for testing (ex: send specific seeds), 0 for real runs
## These next variables are the values passed to the GA
REPRODUCTION=3 ## Number (not fraction!) of individuals formed through reproduction
CROSSOVER=36 ## Number (not fraction!) of individuals formed through crossover
MUTATION=1 ## Probability of mutation (divided by 100)
SIGMA=5 ## Standard deviation for the mutation operation (divided by 100)
ROULETTE=2 ## Percent of individuals selected through roulette (divided by 10)
TOURNAMENT=8 ## Percent of individuals selected through tournament (divided by 10)
RANK=0 ## Percent of individuals selected through rank (divided by 10)
ELITE=0 ## Elite function on/off (1/0)
#####################################################################################################################################################
|
|
Draft
|
Mon Oct 24 17:44:51 2022 |
Ryan Debolt | Icemc inputs | Here is our current assumption of the antenna angles needed for the icemc inputs. |
| Attachment 1: 261D6FA3-5612-412F-80FA-8AD961F7331A.jpeg
| 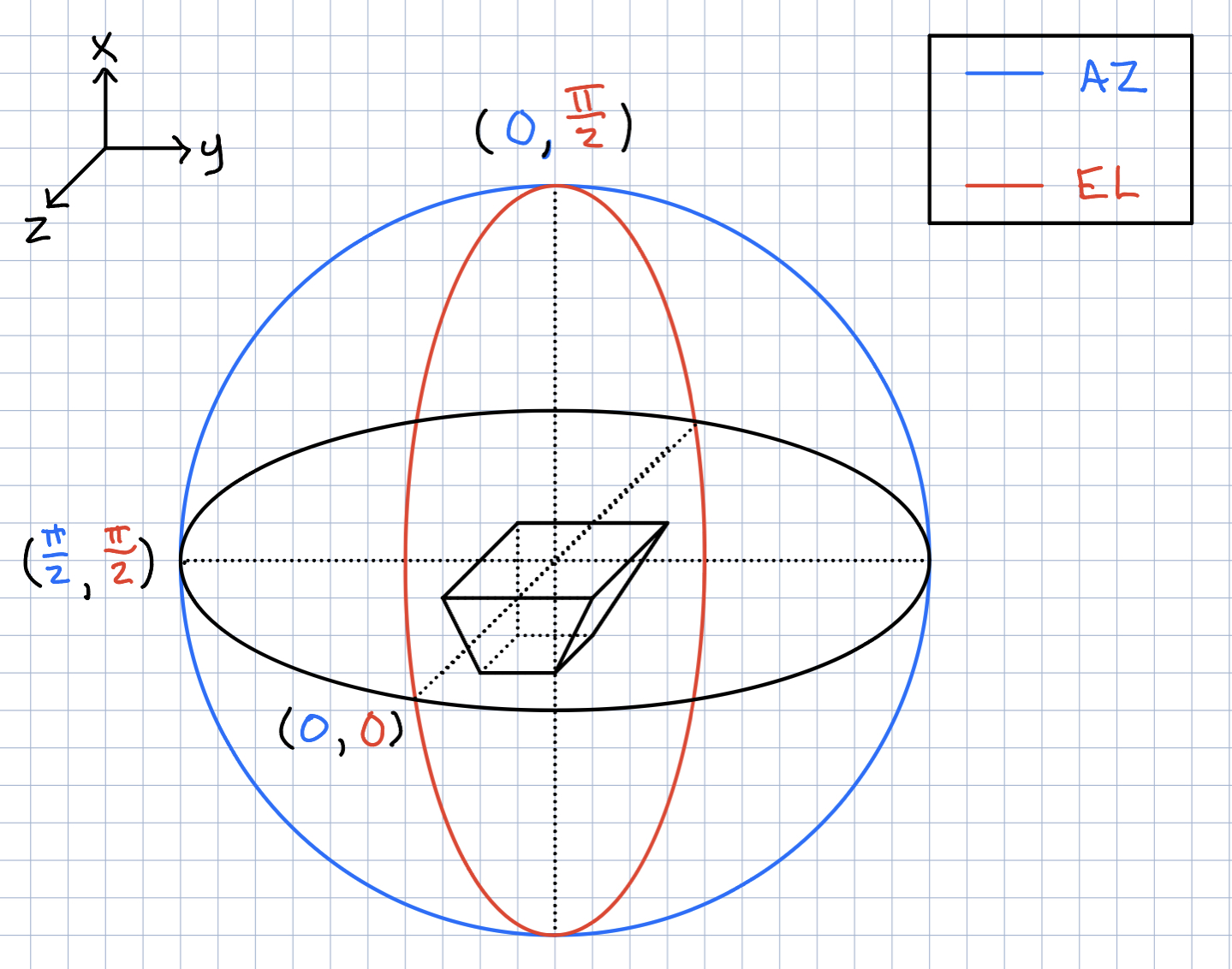 |
|
182
|
Fri Sep 2 12:40:17 2022 |
Alan S | Ice Shell Around Bicone | Here is a PDF with a summary/log of my work trying to integrate an ice shell surrounding the bicone antennas in the GENETIS loop. I made gain patterns of the ARA bicone with the ice shells and without them, showing the impact of the ice on the antenna's gain. Having an ice shell changes these patterns, which are also dependent on the radius of the shell. I showed that convergence on the gain patterns and expected features start ocurring for shells with radii larger than 10 meters. I also keep track of the computation time which increases with bigger shells.
Note: I also add PowerPoint slides that contain all the graphs that I made. The PDF does not contain some of those since I didn't think they were coherent with my write-up, but I may be making an omission.
|
| Attachment 1: XF_ARA_bicone_in_ice_and_air_column.pptx
|
| Attachment 2: Far_Zone_Gain_of_ARA_Bicone_insice_Ice.pdf
|
|
217
|
Wed May 10 17:43:20 2023 |
Alex M | How to run the loop | In this post, I'm going to give step by step instructions on everything you need to do to run the loop. See Julie's thesis for the most recent iteration of this manual. Also refer to entry 123 for a list of some errors we have encountered historically in the loop and how to resolve them, though some fixes may be outdated.
Running the loop
Getting started
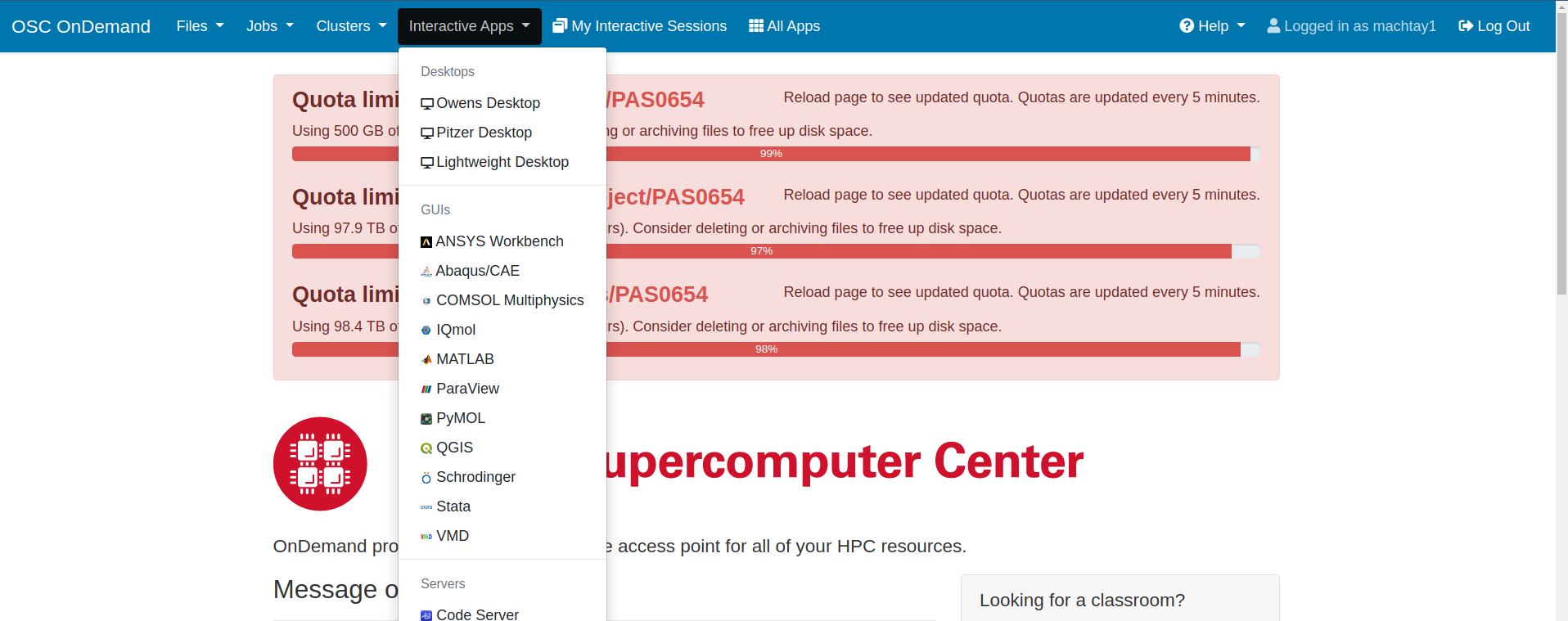
Assuming you have followed the instructions from the onboarding tutorials, you should be ready to access OSC through the interactive website. Instead of using ssh to access OSC, the loop requires that you go to this website: https://ondemand.osc.edu/ . Login with your usual OSC credentials. Once there, you will need to request a Lightweight Desktop by clicking on "Interactive Apps" at the top of the page. See the screenshots attached for how to do this. Old documentation may refer to the Lightweight Desktop as a VDI.
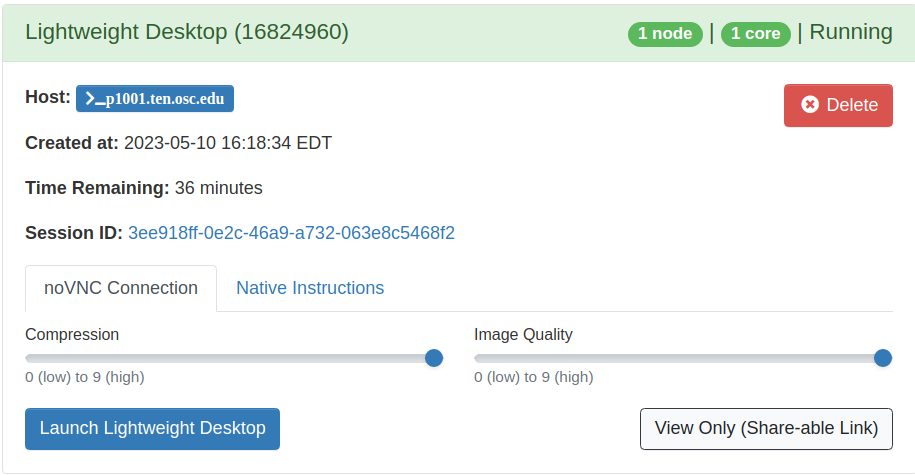
You can request up to 24 hours for a Lightweight Desktop, and it should begin almost immediately. After you request it, the website will take you directly to the waiting page. You can navigate to this page yourself by clicking "My Interactive Sessions" at the top of the ondemand page. When the job begins, you will see a blue buttong that says "Launch Lightweight Desktop". Click it and you should have a new tab open that brings you to a remote desktop environment on OSC that you interact with through your browser.
New runs
If you're starting a new run for the first time (that is, beginning from generation 0), you will need to edit the script that runs the loop. Here is the path: /fs/ess/PAS1960/HornEvolutionOSC/GENETIS_PUEO/BiconeEvolution/current_antenna_evo_build/XF_Loop/Evolutionary_Loop/Loop_Scripts/Asym_XF_Loop.sh . At the top of the script is a list of variables that can be changed. See the first screenshot attached here.
For a new run, you must change the variable <RunName>, which is the first variable in the list. It should be in quotes and the convention is to start the name off with the date of the run, using YYYY/MM/DD format--this will always keep everything in chronological order when listing directories with <ls>. Other variables are explained in the comments next to them in Asym_XF_Loop.sh and you should change them depending on the type of run you are conducting.
To begin the loop, type in the command ./Loop_Scripts/Asym_XF_Loop.sh from the Evolutionary_Loop directory.
When the loop begins, you will be asked to press any key. After you hit a key, the loop will run Part A, which executes the GA using the <start> mode. It will then automatically move on to Part B, which creates the antennas in XFdtd. XF will ask you if you want to specify a location for the XF file to be saved. Click <No>. The loop will automatically save the XF directory into the run directory when it finishes with the first part of part B. In principle (that is, barring errors that stall the loop or interruptions due to the VDI job ending), you will never need to manually enter anything to the loop for it to run beyond this point.
However, if it is your first time using XF, you will need to set the max GHz setting upon startup to 2 instead of the default 10.
Continuing Runs
After starting a new run, the loop will run automatically. When your Lightweight Desktop job ends, you'll need to get another one repeat the steps above. You will not be prompted for any input.
Stepping Back
At times, you may want to continue the loop from an earlier point or interrupt the loop and step back. Usually you'll only want to do this within a generation, but it's possible that you will encounter an error or bug at the end of a generation and wish to return back to that point to remedy it before continuing forward in the next generation. This is a very tricky and delicate process that requires a lot of attention to detail. In the future, we'd like to make an option for the loop to do this automatically when given input at the beginning, but implementing that will take a long time and will be very sophisticated.
The state of the loop is recorded in the savestate file located here: /fs/ess/PAS1960/HornEvolutionOSC/GENETIS_PUEO/BiconeEvolution/current_antenna_evo_build/XF_Loop/Evolutionary_Loop/saveStates . The file will be named <Run_Name>.savestate.txt . In it, you will see three numbers. The first one represents the current generation of the loop. The second one represents the state of the loop, that is, which part in the generation it is currently on. The third one represents the individual it is processing. In practice, the third number is rarely relevant, and is almost always 1, though it can be used for Part B, where the loop is processing individuals sequentially, so if it was interrupted on individual eight (for example) you can change the number to be 8 so that it will pick up where it left off without needing to redo the first seven individuals.
If you want to step back to an earlier state in a generation, you'll need to interrupt the loop and revert the state by changing the second number in the savestate file. This is NOT all you will have to do, however. Because of the way files are edited and moved around during states, you will need to manually revert files to be in the format or location needed by the earlier state.
For example, you may need to revert from state 2 (when XF is creating the antenna geometries) to state 1 (when the GA generates new individuals). In this case, you'll need to edit the savestate file to change the second number from a 2 to a 1. Then, you'll need to change the generationDNA.csv file in Generation_Data to use the values from the previous generation, since the GA will have overwritten that file and uses it to determine what individuals are to be selected from. To do this, you'll need to locate the <gen>_generationDNA.csv file from the previous generation in the <RunName> directory in Run_Outputs. This is should be found here: /fs/ess/PAS1960/HornEvolutionOSC/GENETIS_PUEO/BiconeEvolution/current_antenna_evo_build/XF_Loop/Evolutionary_Loop/Run_Outputs/<RunName>/Generation_Data .
The above is a relatively simple example of how to step back in the loop. Stepping back from a late state to an early state will be more complicated, though if you're willing to sacrifice run time you can pretty easily follow the above to just restart a generation. Stepping back to a previous generation will be explained in a future update to this post. |
| Attachment 1: Ondemand_Front_Page.png
|  |
| Attachment 2: Ondemand_Interactive_Apps.png
|  |
| Attachment 3: Launching_Lightweight_Desktop.png
|  |
|
177
|
Tue Aug 9 15:28:50 2022 |
Ryan Debolt | How many individuals to use in the GA. | One of our foundational questions tied to the optimization of the GA has been "How many individuals should we simulate". Up to now, our minds were made up for us by the speed of arasim being great enough that the time cost of simulating individuals was great enough that the improvements made from having more were not enough to justify the slowdown. However, with the upgrade to the faster, more recent version of arasim, I decided to re-examine this. This was also spurred on by the fact that the last time I ran this test we were testing GA performance by final generation metrics rather than by how many generations it took to reach a benchmark. So in one of my optimization tests, I tracked this data.
To start, using the same run proportions, using a .5 chi-squared benchmark, the average time across all 89 run types used in this run was 25.4 generations for 50 individuals as compared to 8.3 generations running the same test for 100. Furthermore, the minimum number of generations for 50 individuals was 4.8 while using 100 individuals yielded 2.4. So on average running 100 individuals was about 3 times fast at reaching this benchmark than with 50. And when comparing the best result regardless of run type, 100 individuals was still 2 times quicker than the min for 50 individuals. Finally, the run that yielded an average of 2.4 generations for 100 individuals took an average of 29.2 generations with 50 individuals or roughly 12 times the generations.
For the test we will discuss, we ran 89 different run types that all used 60% rank, 20% roulette, and 20% tournament selection respectively. These test had the following ranges:
6-18% of individuals through reproduction (steps of 3%)
64-88% of individuals through crossover (steps of 12%)
0-10% mutation rate (steps of 5%)
1-5% sigma on mutation (steps of 1%)
These tests also used our fitness scores with simulated error of .1 to imitate arasim's behavior and as such we used the chi-squared value to evaluate these scores as there is no error on those values.
Comparing this same test with a tighter chi-squared benchmark of .25, we see similar results. On average 50 individuals took 37.1 generations to reach this point while 100 individuals took 16.0 generations. Similarly, the minimums amount of gens for 50 individuals was 15.4 while 100 individuals was 5. Finally, the corresponding run for the 5 generation min with 100 individuals took 41.8 generations with 50 individuals. These correspond to speed up's of 2.3, 3.08, and 8.36 respectively.
This data implies that on average, independent of run type, we should expect to have to use 2-3 times fewer generations while running 100 individuals than we would running 50 individuals but we could see up to 8-12 times fewer generations to reach benchmarks. Another data set using a different set of selection methods was also tested for this and again yielded similar results, though overall the runs from the first batch were better across both 50 and 100 individuals and so those results are likely to be more indicative of the parameters we use in a true run.
The data being examined in these results can be found here: https://docs.google.com/spreadsheets/d/1GlfnjQSO6VI8MuUGYTUcLkjwDZU98nyFFysgTTfVFOE/edit?usp=sharing |
|
Draft
|
Wed Mar 25 21:22:00 2026 |
Ryan Debolt | How many individuals to use in the GA. |
| Quote: |
|
One of our foundational questions tied to the optimization of the GA has been "How many individuals should we simulate". Up to now, our minds were made up for us by the speed of arasim being great enough that the time cost of simulating individuals was great enough that the improvements made from having more were not enough to justify the slowdown. However, with the upgrade to the faster, more recent version of arasim, I decided to re-examine this. This was also spurred on by the fact that the last time I ran this test we were testing GA performance by final generation metrics rather than by how many generations it took to reach a benchmark. So in one of my optimization tests, I tracked this data.
To start, using the same run proportions, using a .5 chi-squared benchmark, the average time across all 89 run types used in this run was 25.4 generations for 50 individuals as compared to 8.3 generations running the same test for 100. Furthermore, the minimum number of generations for 50 individuals was 4.8 while using 100 individuals yielded 2.4. So on average running 100 individuals was about 3 times fast at reaching this benchmark than with 50. And when comparing the best result regardless of run type, 100 individuals was still 2 times quicker than the min for 50 individuals. Finally, the run that yielded an average of 2.4 generations for 100 individuals took an average of 29.2 generations with 50 individuals or roughly 12 times the generations.
For the test we will discuss, we ran 89 different run types that all used 60% rank, 20% roulette, and 20% tournament selection respectively. These test had the following ranges:
6-18% of individuals through reproduction (steps of 3%)
64-88% of individuals through crossover (steps of 12%)
0-10% mutation rate (steps of 5%)
1-5% sigma on mutation (steps of 1%)
These tests also used our fitness scores with simulated error of .1 to imitate arasim's behavior and as such we used the chi-squared value to evaluate these scores as there is no error on those values.
Comparing this same test with a tighter chi-squared benchmark of .25, we see similar results. On average 50 individuals took 37.1 generations to reach this point while 100 individuals took 16.0 generations. Similarly, the minimums amount of gens for 50 individuals was 15.4 while 100 individuals was 5. Finally, the corresponding run for the 5 generation min with 100 individuals took 41.8 generations with 50 individuals. These correspond to speed up's of 2.3, 3.08, and 8.36 respectively.
This data implies that on average, independent of run type, we should expect to have to use 2-3 times fewer generations while running 100 individuals than we would running 50 individuals but we could see up to 8-12 times fewer generations to reach benchmarks. Another data set using a different set of selection methods was also tested for this and again yielded similar results, though overall the runs from the first batch were better across both 50 and 100 individuals and so those results are likely to be more indicative of the parameters we use in a true run.
The data being examined in these results can be found here: https://docs.google.com/spreadsheets/d/1GlfnjQSO6VI8MuUGYTUcLkjwDZU98nyFFysgTTfVFOE/edit?usp=sharing
|
|
|
210
|
Wed Apr 5 17:21:07 2023 |
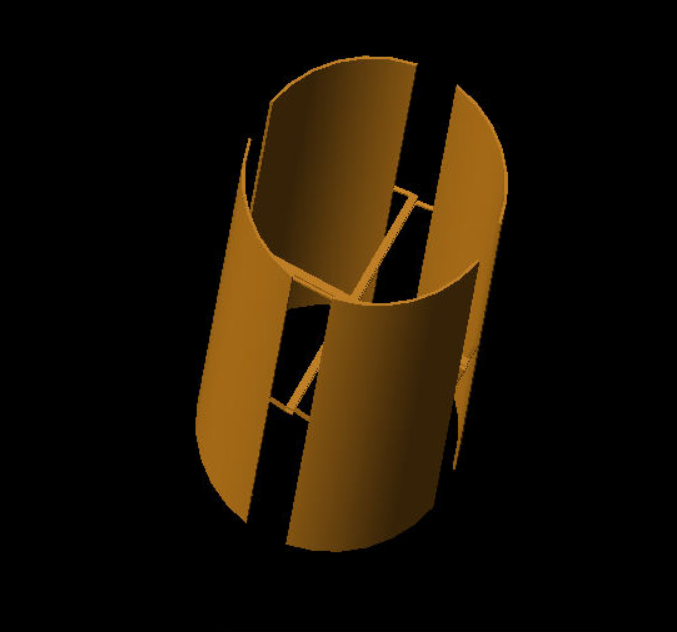
Nick King | H-pol XF Design | Work in progress. Update on H-pol XF design. The first plot is the gain pattern with ferrite rod sets (blue) vs. with nothing but the copper plates.
The second plot is the gain pattern with everything but the ferrite rod sets vs. with nothing but the copper plates.
The ferrite rod sets function to narrow the gain pattern. |
| Attachment 1: Screenshot_(7).png
| .png) |
| Attachment 2: Screenshot_(8).png
| .png) |
| Attachment 3: HpolfromScript(5_10_2023).png
| .png) |
| Attachment 4: StephHpolwith_just_copper_plates_and_wires.png
|  |
|
222
|
Thu Jun 1 21:49:29 2023 |
Dylan Wells | Guide to Updating pueoSim | How To Update PueoSim For GENETIS:
First, whoever updates pueoSim needs access to pueoBuilder, pueoSim, and niceMC on GitHub (ask Will for permissions).
Once you do, go into the peuoSim directory at /fs/ess/PAS1960/buildingPueoSim/
and source set_env.sh
`source set_env.sh`
Then, we want to make copies of the files we are currently modifying for the GENETIS loop.
For PueoSim v1.1.0 these are:
-
nicemc/src/EventGenerator.cc # Modified to create custom output root files to calculate errors
-
pueosim/test/simulatePueo.cc # Modified to stop the simulation of the LF antenna which is not needed for GENETIS
-
pueosim/src/Seavey.cc # Modified to read in custom gain files
-
pueosim/src/pueo.cc # Modified to read in custom gain files
-
pueosim/include/Seavey.hh # Modifies to read in custom gain files
Currently, I store copies of these in the `/fs/ess/PAS1960/buildingPueoSim/backups` directory in case somebody accidentally overwrites the files in pueoBuilder.
Once you’ve made the copies, you can run `./pueoBuilder.sh` from the `/fs/ess/PAS1960/buildingPueoSim/pueoBuiler` directory. This will rebuild pueoSim and niceMC, pulling the latest updates from GitHub.
You may need to delete the pueoSim and niceMC directories in order for the builder to pull the latest version from GitHub. Or, if it’s being really stubborn, move the whole pueoBuilder directory to a temporary location and run the builder from scratch with
`git clone git@github.com:PUEOCollaboration/pueoBuilder` and then `./pueoBuilder.sh`
Then, you will need to reference the copies of the changed files to make changes to the new version of pueoSim. Hope this doesn’t cause too much of a headache, and when you’re done, return to the /fs/ess/PAS1960/buildingPueoSim/pueoBuiler directory.
Then you simply type
`make install`
then
`make`
And now, pueoSim should be ready to run!
EventGeneratior.cc Changes and Rationale:
PueoSim’s default output ROOT files are very large and therefore time-consuming to parse through to get the information we need to calculate effective volume and errors. So, we want to create a custom ROOT file with only the variables we want, greatly increasing the speed of the analysis.
To do this, we want to create a new TFile with corresponding TTree and TBranches that will store the loop.positionWeight, loop.directionWeight, and neutrino.path.weight. Then, we want to fill the Tree when a detector is triggered and write the results to the file at the end of the loop.
Sample code for initializing the TFile:

simulatePueo.cc Changes and Rationale:
By default, pueoSim v1.1.0 runs a simulation for the normal antenna and a low-frequency (LF) antenna. As GENETIS is evolving for the main antenna, we are not interested in using computation time to simulate the LF antenna. So, we comment out the lines that initialize the LF detector in this file.
Seavey.cc, Seavey.hh, and pueo.cc Changes and Rationale:
As of pueoSim v1.1.0, the simulation software only has the ability to read in one antenna. This is a problem, as we want it to be able to read in files for any antenna we make. So, we need to change these scripts to be able to read in whatever gain files we want.
The convention the loop uses is to input gain files in the format of
vv_0_Toyon{runNum}
hh_0_Toyon{runNum}
…
In the ./pueoBuilder/components/pueosim/data/antennas/simulated directory
So, the main change is getting the runNum variable into Seavey.cc file where the gains are read. Currently, we define a global variable at the stary of pueo.cc and pass it into where Seavey is called. This means we have to add runNum as a parameter in Seavey.cc as well as Seavey.hh. Finally, we change the name of the read-in files to be in the above format AFTER dividing runNum by 1000 (this is because pueoSim uses the run number as a random number generating seed, so we divide runNum by 1000 to be able to read in the same gain patterns for multiple seeds of the same individual).
Note: We used to change pueoSim to output a veff.csv file. We now handle this calculation by analyzing ROOT files, so this change is no longer necessary. |
|
Draft
|
Sun Mar 29 08:55:02 2026 |
Dylan Wells | Guide to Updating pueoSim | How To Update PueoSim For GENETIS:
First, whoever updates pueoSim needs access to pueoBuilder, pueoSim, and niceMC on GitHub (ask Will for permissions).
Once you do, go into the peuoSim directory at /fs/ess/PAS1960/buildingPueoSim/
and source set_env.sh
`source set_env.sh`
Then, we want to make copies of the files we are currently modifying for the GENETIS loop.
For PueoSim v1.1.0 these are:
-
nicemc/src/EventGenerator.cc # Modified to create custom output root files to calculate errors
-
pueosim/test/simulatePueo.cc # Modified to stop the simulation of the LF antenna which is not needed for GENETIS
-
pueosim/src/Seavey.cc # Modified to read in custom gain files
-
pueosim/src/pueo.cc # Modified to read in custom gain files
-
pueosim/include/Seavey.hh # Modifies to read in custom gain files
Currently, I store copies of these in the `/fs/ess/PAS1960/buildingPueoSim/backups` directory in case somebody accidentally overwrites the files in pueoBuilder.
Once you’ve made the copies, you can run `./pueoBuilder.sh` from the `/fs/ess/PAS1960/buildingPueoSim/pueoBuiler` directory. This will rebuild pueoSim and niceMC, pulling the latest updates from GitHub.
You may need to delete the pueoSim and niceMC directories in order for the builder to pull the latest version from GitHub. Or, if it’s being really stubborn, move the whole pueoBuilder directory to a temporary location and run the builder from scratch with
`git clone git@github.com:PUEOCollaboration/pueoBuilder` and then `./pueoBuilder.sh`
Then, you will need to reference the copies of the changed files to make changes to the new version of pueoSim. Hope this doesn’t cause too much of a headache, and when you’re done, return to the /fs/ess/PAS1960/buildingPueoSim/pueoBuiler directory.
Then you simply type
`make install`
then
`make`
And now, pueoSim should be ready to run!
EventGeneratior.cc Changes and Rationale:
PueoSim’s default output ROOT files are very large and therefore time-consuming to parse through to get the information we need to calculate effective volume and errors. So, we want to create a custom ROOT file with only the variables we want, greatly increasing the speed of the analysis.
To do this, we want to create a new TFile with corresponding TTree and TBranches that will store the loop.positionWeight, loop.directionWeight, and neutrino.path.weight. Then, we want to fill the Tree when a detector is triggered and write the results to the file at the end of the loop.
Sample code for initializing the TFile:

simulatePueo.cc Changes and Rationale:
By default, pueoSim v1.1.0 runs a simulation for the normal antenna and a low-frequency (LF) antenna. As GENETIS is evolving for the main antenna, we are not interested in using computation time to simulate the LF antenna. So, we comment out the lines that initialize the LF detector in this file.
Seavey.cc, Seavey.hh, and pueo.cc Changes and Rationale:
As of pueoSim v1.1.0, the simulation software only has the ability to read in one antenna. This is a problem, as we want it to be able to read in files for any antenna we make. So, we need to change these scripts to be able to read in whatever gain files we want.
The convention the loop uses is to input gain files in the format of
vv_0_Toyon{runNum}
hh_0_Toyon{runNum}
…
In the ./pueoBuilder/components/pueosim/data/antennas/simulated directory
So, the main change is getting the runNum variable into Seavey.cc file where the gains are read. Currently, we define a global variable at the stary of pueo.cc and pass it into where Seavey is called. This means we have to add runNum as a parameter in Seavey.cc as well as Seavey.hh. Finally, we change the name of the read-in files to be in the above format AFTER dividing runNum by 1000 (this is because pueoSim uses the run number as a random number generating seed, so we divide runNum by 1000 to be able to read in the same gain patterns for multiple seeds of the same individual).
Note: We used to change pueoSim to output a veff.csv file. We now handle this calculation by analyzing ROOT files, so this change is no longer necessary. |
|
123
|
Wed Dec 2 15:24:39 2020 |
Alex M | Guide for Loop Errors | Attached is a .txt file you can find in the Evolutionary_Loop directory as Loop_Error_List.txt. It's a list of the current errors we sometimes experience in the loop, along with how to fix them if you encounter them while running. If people encounter errors that aren't in the list, let everyone know in the #genstudents chat on slack and update the file with the error message and when it was encountered (what state the loop was in) and the possible cause and solution if you know it. |
| Attachment 1: Loop_Error_list.txt
|
Below is a list of errors we may encounter in the loop as of 11/25/20:
Error: "Pre-while, pre-for"
Description: This is an error you'll encounter after AraSim has "completed." The loop will hang after outputting "Pre-while" and then "Pre-for." This comes from the fitness function--the loop is indicating that it is inside the fitness function right before it enters the two loops it runs over the AraSim data. Hanging here indicates that there was an issue running AraSim. Specifically, it indicates that at least one of the jobs for the *first* individual in AraSim failed.
Potential causes:
This may be caused by an issue in generating the gain files used for AraSim. These gain files are placed in the AraSim directory under the names a_{num}.txt, where {num} represents the individual number. You can check a_1.txt in the AraSim directory and see if it's complete (if it isn't, you can usually tell just by opening the file and seeing that only two lines have been printed to it).
One way of looking for the cause of this error is to look at the job error files. Inside the {runname} directory are directories containing the error and output files from the AraSim jobs. These are .../{gen}_AraSim_Errors and .../{gen}_AraSim_Output, where {gen} is the generation number. One example of an error message I've seen in the error files was
"
terminate called after throwing an instance of 'std::out_of_range'
what(): basic_string::substr
/var/spool/slurmd/job2345909/slurm_script: line 37: 171196 Aborted (core dumped) ./AraSim setup.txt $runNum outputs/ a_${num}.txt > $TMPDIR/AraOut_${gen}_${num}_${Seeds}.txt
"
This appeared in all of the error files for the first individual's jobs.
Resolution:
The best way to resolve this is to start by checking the error files. In the case of the error message above, it would be best to go to the AraSim directory and check for the a_{num}.txt file. If you see just one (ex: a_1.txt), then that's likely to be the culprit (especially if the file is obviously not completely filled)--these files should be removed, so if one is left it may not have been moved correctly, likely due to permissions errors. Remove the a_{num}.txt file and restart from the AraSim job submissions (potential speed up: add part of the error message to the self-correcting phrases in Part_D2_AraSeed.sh to only rerun those individuals).
It's also possible that the issue was caused in XF. Make sure you follow the instructions above and start back at stage 2 to restart from the beginning of XF if starting back from AraSim doesn't work. Try to take notes on the differences you see to add to this.
It's also notable that this may be caused by permissions issues. Every time someone is handing the loop off to someone else, the OpenPermissions.sh script should be run (passing the {runName} as an argument). Look in that script to determine which files need to have open permissions. If the person with ownership of the closed files isn't available to open them, you can remove them and start back from where they would have been created. This usually occurs in AraSim and the AraOut files in Antenna_Performance_Metric should have their permissions fixed or be removed (for that generation only).
*****************************************************************************
Error: <Loop hangs while outputting dimensions and fitness scores>
Description:
This is similar to the error above, except that instead of hanging on the first individual, it hangs on some later individual.
Resolution:
The instructions for resolving this should be the same as the ones above. This seems to be less common and is usually resolved by the self-correcting code in Part_D2. Regardless, if you encounter this error the first step should be to follow the instructions below in case there is just one or a handful of failed AraSim jobs. If that doesn't work, step back to stage 5 to resubmit the AraSim jobs after clearing out the possible offending files. If that doesn't work, step back to XF (you can always just step back to XF at the beginning if you're unsure that stepping back to AraSim will resolve this to potentially save time).
It's also possible that there is an error in just a handful (or even just one) of the AraSim jobs. This might be caused by opening permissions after someone has already taken over running the loop. In this case, you might be able to start the loop back up without needing to resubmit all of the AraSim jobs or step back all the way to XF. To do this, you'll need to figure out which AraSim job failed. Check the AraSim error files and output files for that generation (specifically, check to see if one is *missing*). You should be able to figure out which individual the loop is stuck on by counting how many sets of dimensions and fitness scores were printed to screen before the loop started hanging. Go to /Antenna_Performance_Metric (inside .../Evolutionary_Loop) and list all of the AraOut files corresponding to that individual and check them to see if any of them appear incomplete (AKA don't have an effective volume at the bottom).
Once you find the individual jobs that failed, you can set up the loop to only rerun those jobs. First, go to the AraSim flags directory inside the RunName directory and populate the the flags like so:
>for i in `seq 1 <NPOP>
>do
>for j in `seq 1 <jobs per individual>
>do
>echo <gen> > ${i}_${j}.txt
>echo $i >> ${i}_${j}.txt
>echo $j >> ${i}_${j}.txt
>done
>done
This will populate all of the flag files needed for AraSim to move on. Remove the flag files corresponding to the identified failed AraSim jobs. Next, go into the AraSim error file directory in the RunName directory (/<gen>_AraSim_Errors) and replace any text inside the error files corresponding to the failed AraSim jobs with the phrase "segmentation violation" (spelling and capitalization matter!). This is one of the phrases used in the self correcting part of the loop in Part_D2 and indicates to the loop to resubmit the AraSim job for that individual.
After doing this, you should be able to return to .../Evolutionary_Loop and change the savestate in /savestates to 6 from 7. Now you can start back the loop and it will tell you that it's waiting for the AraSim jobs to finish. After 1/2 minutes it will notice that the error files have "segmentation violation" in them and will resubmit only the AraSim jobs you specified as having failed.
*****************************************************************************
Error: "cannot connect to X11 forwarding" (or something to that effect)
Description:
This usually occurs during XF, but it may occur during the display of plots. In the case of plots being unable to display, the loop should still be able to operate, though plots might not update. However, if this message occurs during XF the data for the gain patterns for the antennas won't be properly created. This can occur at the first opening of the XF GUI or on the second part (after the xfsolver jobs have run).
Potential causes:
First, you should make sure that you are logged in to OSC using <ssh -XY userID>. The -XY allows x11 forwarding, which is needed for the XF GUI to appear. Also remember to indicate that you need X11 forwarding when requesting your interactive job (using --x11). It's also possible that your connection to the X11 forwarding can be interrupted after a long time (I've seen the loop work for several generations over multiple interactive job submissions and then suddenly get this error).
Resolution:
My advice is to log out and back in to OSC each time your interactive job ends. This is an uncommon error but it's easy to miss. Once you've logged back in, you'll need to restart the save state back to the part of XF where you got this error (either 2 or 3 depending on which par the error appeared in).
*****************************************************************************
Error:
|
|
224
|
Mon Jun 5 15:00:07 2023 |
Ryan Debolt | Github tests | We want to automate tests in pull requests in GitHub to run unit tests on individual functions to ensure they return the expected results. This will help prevent bugged code from being pushed to working branches. We will add a list of unit tests to this ELOG as we design them.
|
|
228
|
Thu Jun 29 12:05:45 2023 |
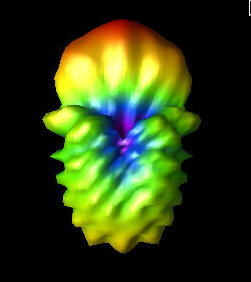
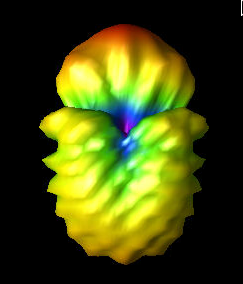
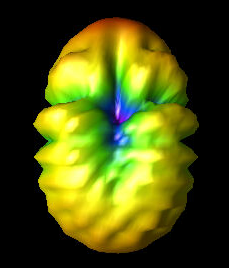
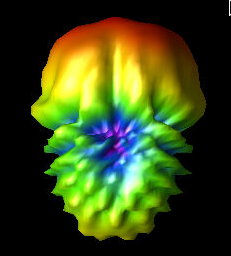
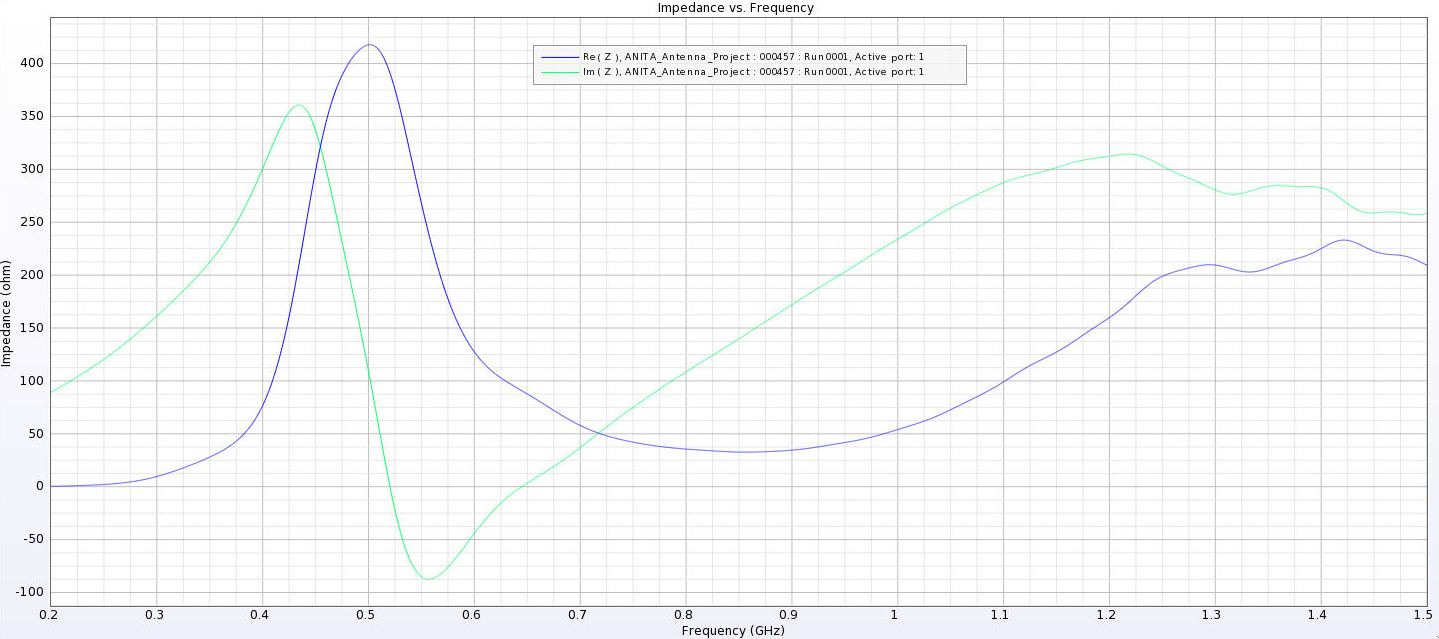
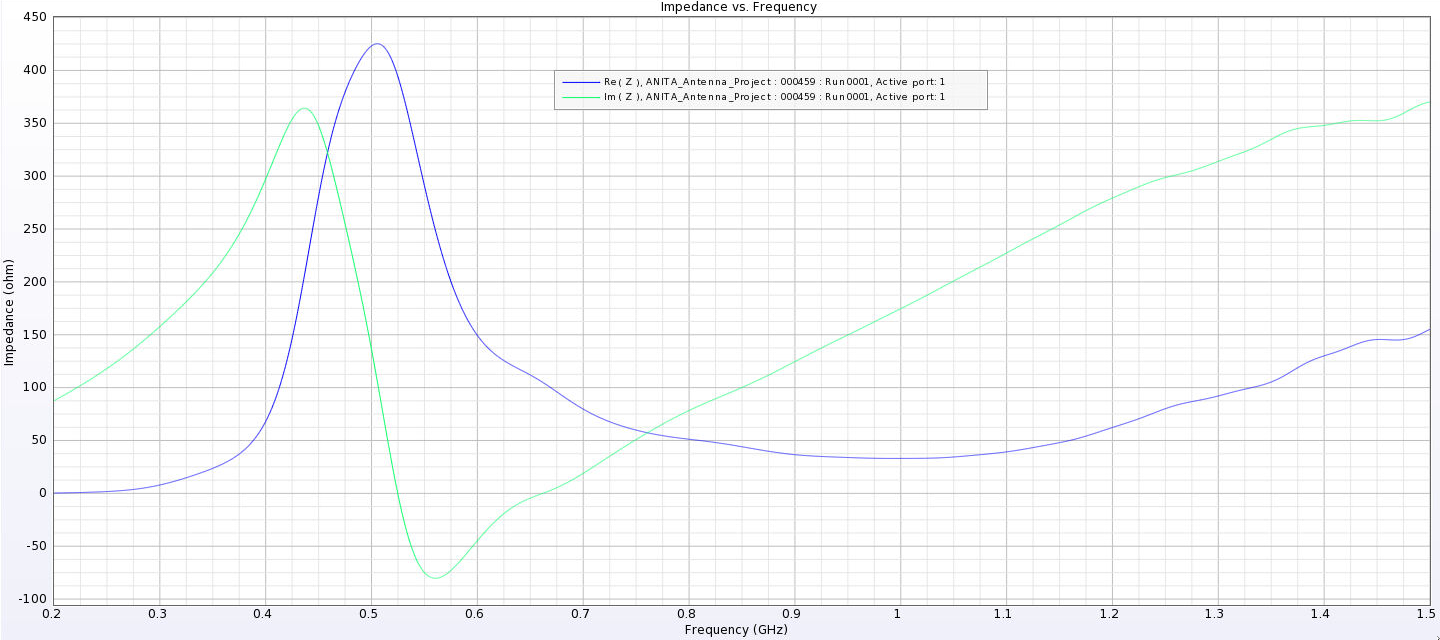
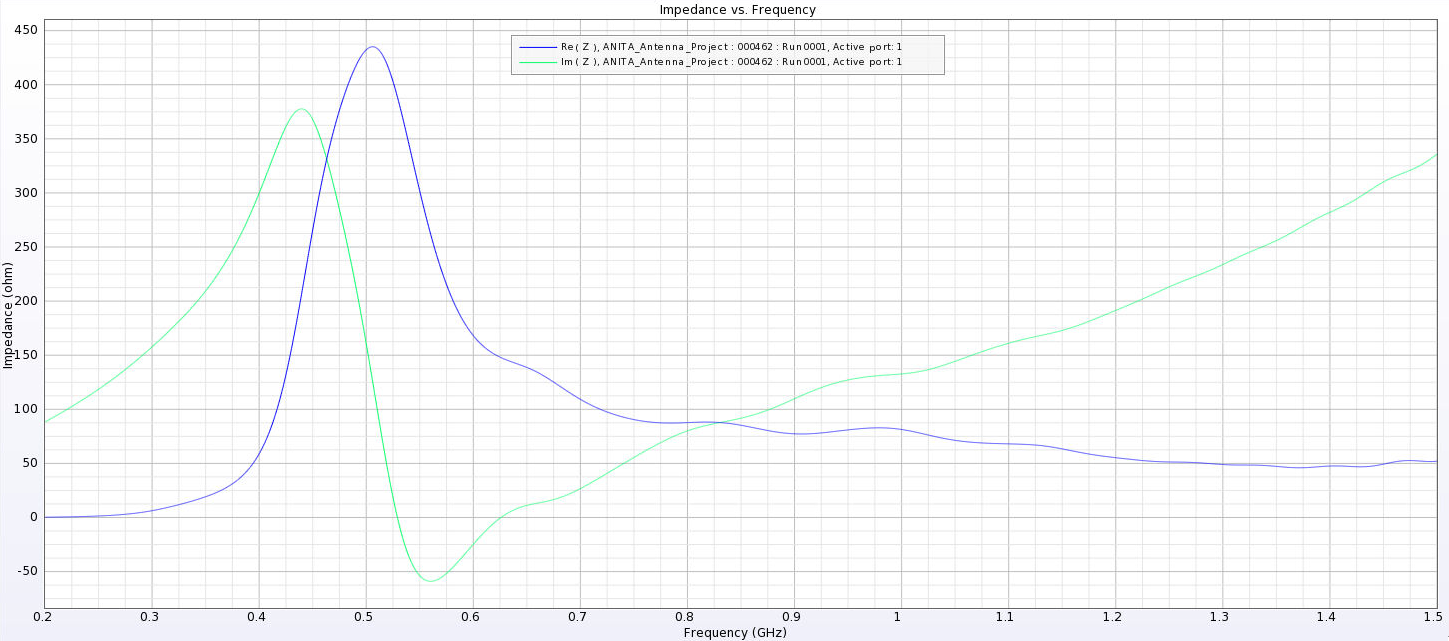
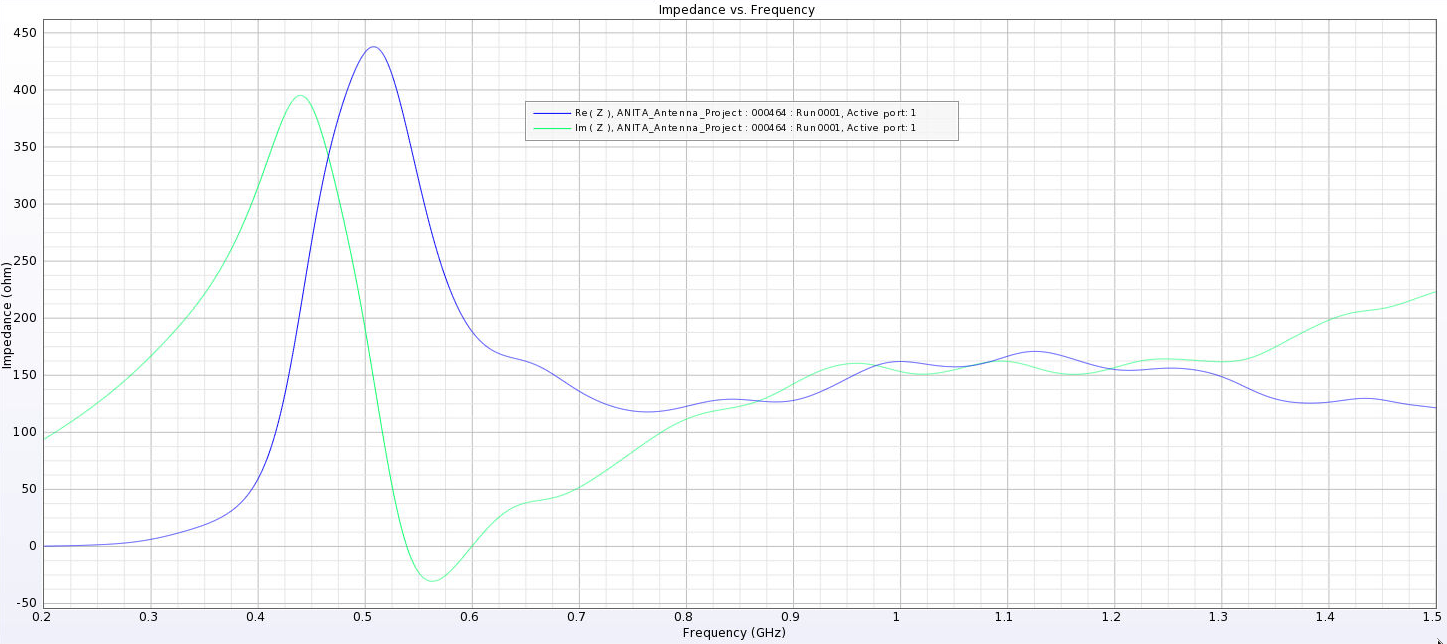
Alex M | Gain at various source heights | Here are a collection of antenna gain patterns with the antenna power source located at various heights at 500 MHz. The heights are indicated in the name of the images and were located at 3cm, 6cm, 12cm, and 24cm from the base. One thing to note is that the power source becomes longer as a function of height (since it's connected to the ridges, which slope back).
The 3D gain plots attached show the gain becoming more uniform until we get to the last one, where it seems to reinvert and be much stronger in the upward direction than below. I'll update this post with some polar slices for a more complete comparison between them.
EDIT: 07/17/23
I've made a few adjustments and added plots of them.
- First, I separated the ridges. Previously, the ridges touched at their inner corners. This electrically connected them and may be why we saw very high peaks in the first set of impedance plots. You can see the plots named Separated_Impedance_<x>_cm.png.
- This appears to have lowered the peaks considerably and also shifted them to lower frequencies. The peaks are still high when the source is placed high.
- We seem to be developing another peak at higher frequencies, although this also existed in the previous tests for the high sources.
- After this, I added a waveguide and positioned the source between the extensions of the ridges into the waveguide. I've also added a picture that shows the waveguide in the geometry.
- The sources here are positioned from 1 cm below the ridges to 8 cm below the ridges. The naming convention is the same as above, except it includes "Waveguide" in the name.
- The peaks are considerably lower here and seem consistent as we place sources further down. They are all below 500 Ohms, compared to as much as 4000 Ohms in the previous tests.
EDIT: 08/10/23
I've been trying more tweaks to the design to get rid of the peaks we see in the plots below. I'm having trouble actually getting the impedance to match close to the 50 Ohm value we expect from the cable. The best case situations seem to come from when the ridges are still touching, which we don't want. Here are the properties I've been changing:
- Ridge positions (gene in GA)
- We don't want the ridges to touch, but I can't get them to be very far from each other anyway. The minimum wavelength in our bandwidth would be 20 cm, but that's close to the size of the bottom of the antenna for when we make it large, so the separation is still small.
- When the ridges do touch is actually when I get the best matching,
- Bottom size (gene in GA)
- I've been changing the size of the bottom of the antenna. This is something that can change substantially between individuals in the GA, so even if I find a good value for this, the impedance it gives wouldn't be the same for all individuals we see in an evolution.
- This does seem to move the location of the impedance peak (in frequency) but I can't seem to get it fully out of band.
- Waveguide length
- I've been changing the length of the waveguide between 3 cm and 10 cm. We said that somewhere in the middle should be the most reasonable (the example antenna we saw looked to have a waveguide of ~5 cm long). It seems like the source position is more important.
- Source position
- I've been moving the position of the power source inside the waveguide. This seems to move around the peak and can lower its height, but I haven't been able to get peaks lower than ~400 Ohms, except in cases where the ridges are in contact.
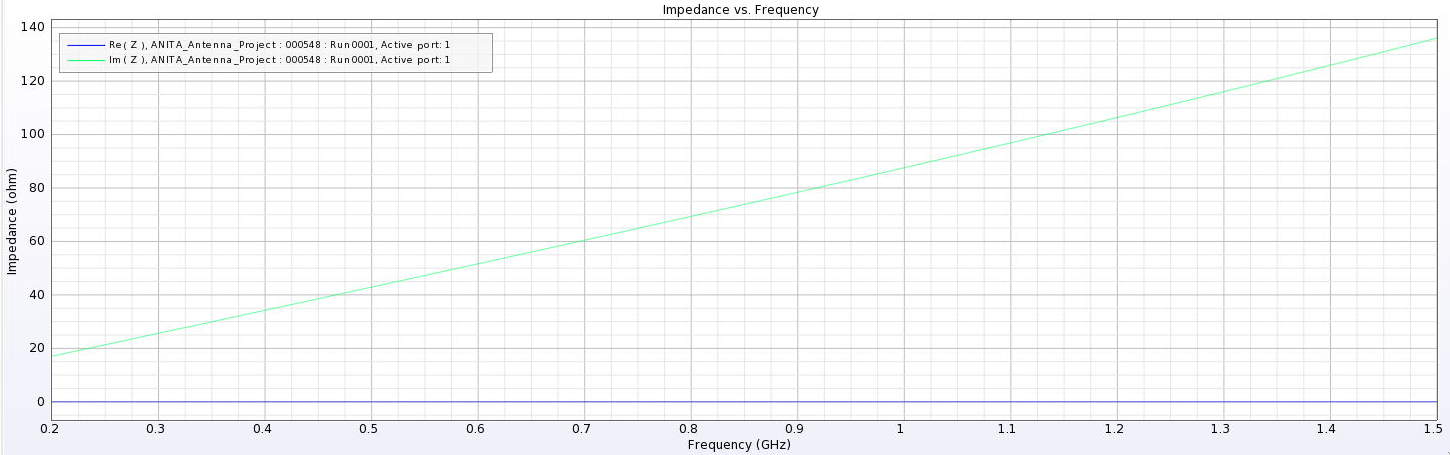
I've added a picture of the best impedance I managed to get, but this involved an antenna with a small base and ridges touching. My next step is to just try a grid search by making many antennas automatically and simulating them all to look for any that have a nice impedance in our band. |
| Attachment 1: 3_cm.png
|  |
| Attachment 2: 6_cm.png
|  |
| Attachment 3: 12_cm.png
|  |
| Attachment 4: 24_cm.png
|  |
| Attachment 5: PUEO_Impedance_3_cm.png
| 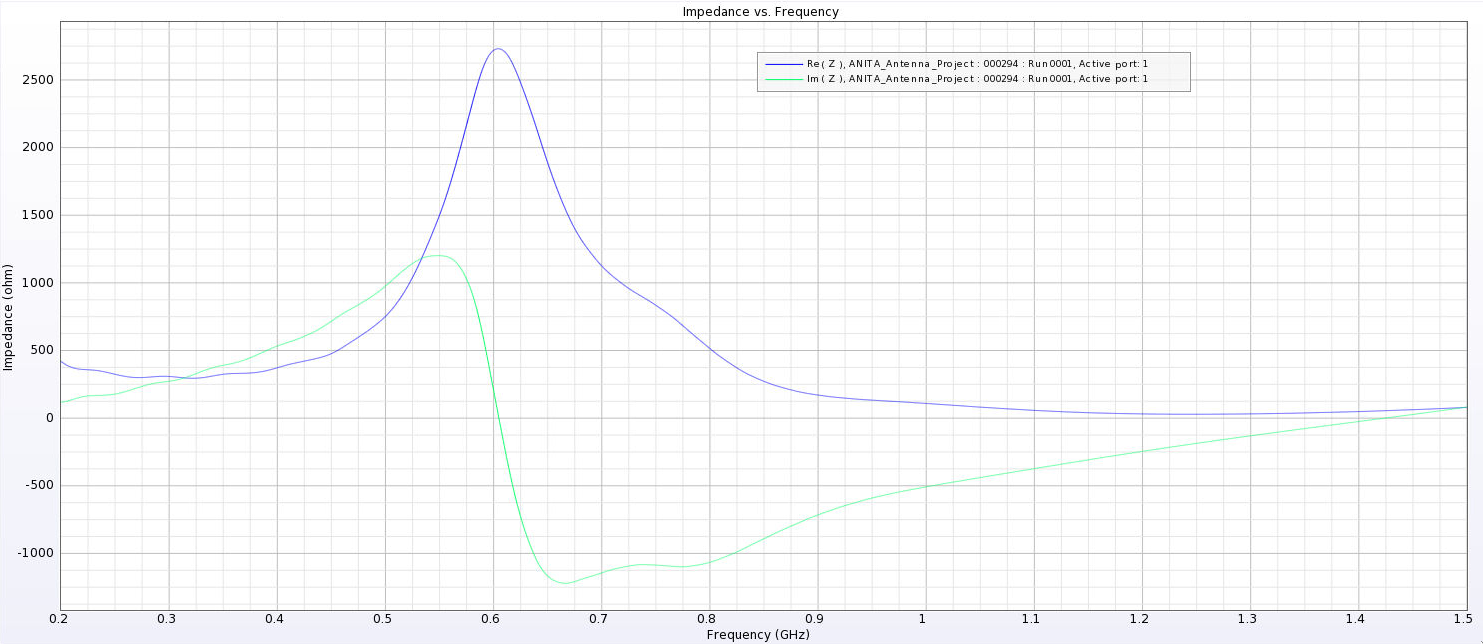 |
| Attachment 6: PUEO_Impedance_6_cm.png
| 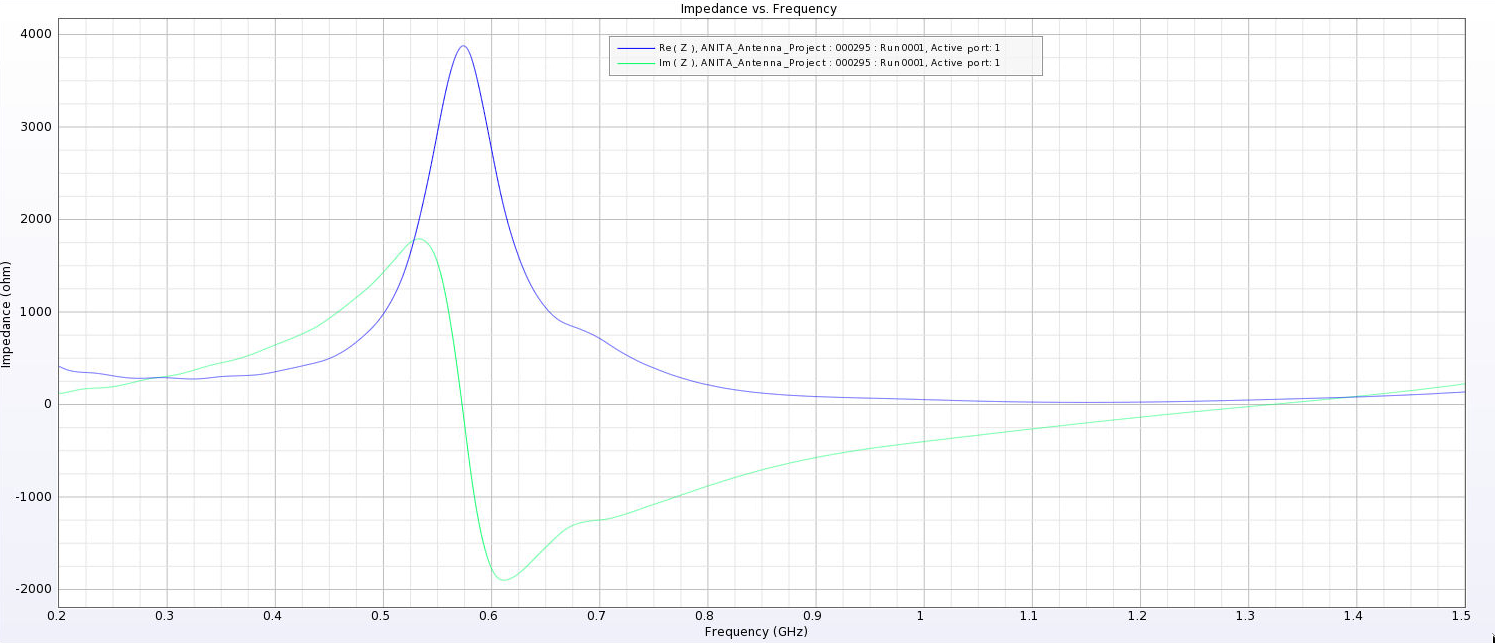 |
| Attachment 7: PUEO_Impedance_12_cm.png
| 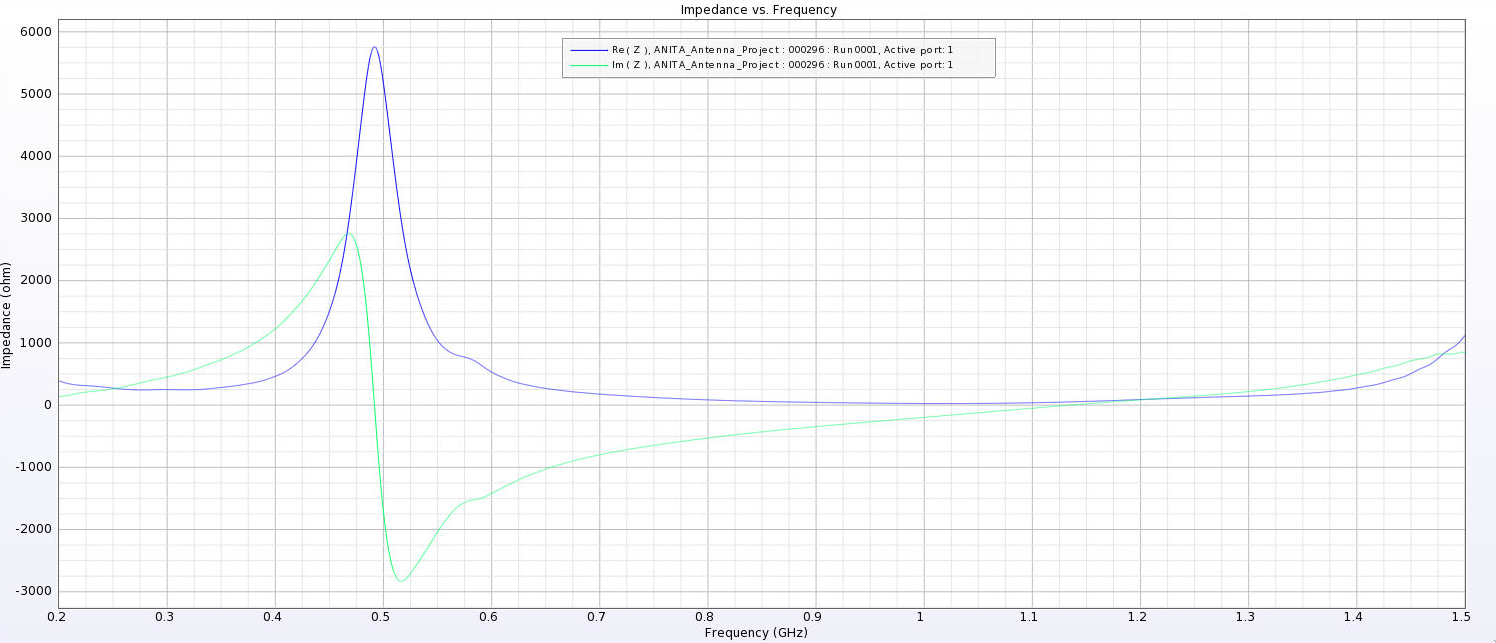 |
| Attachment 8: PUEO_Impedance_24_cm.png
| 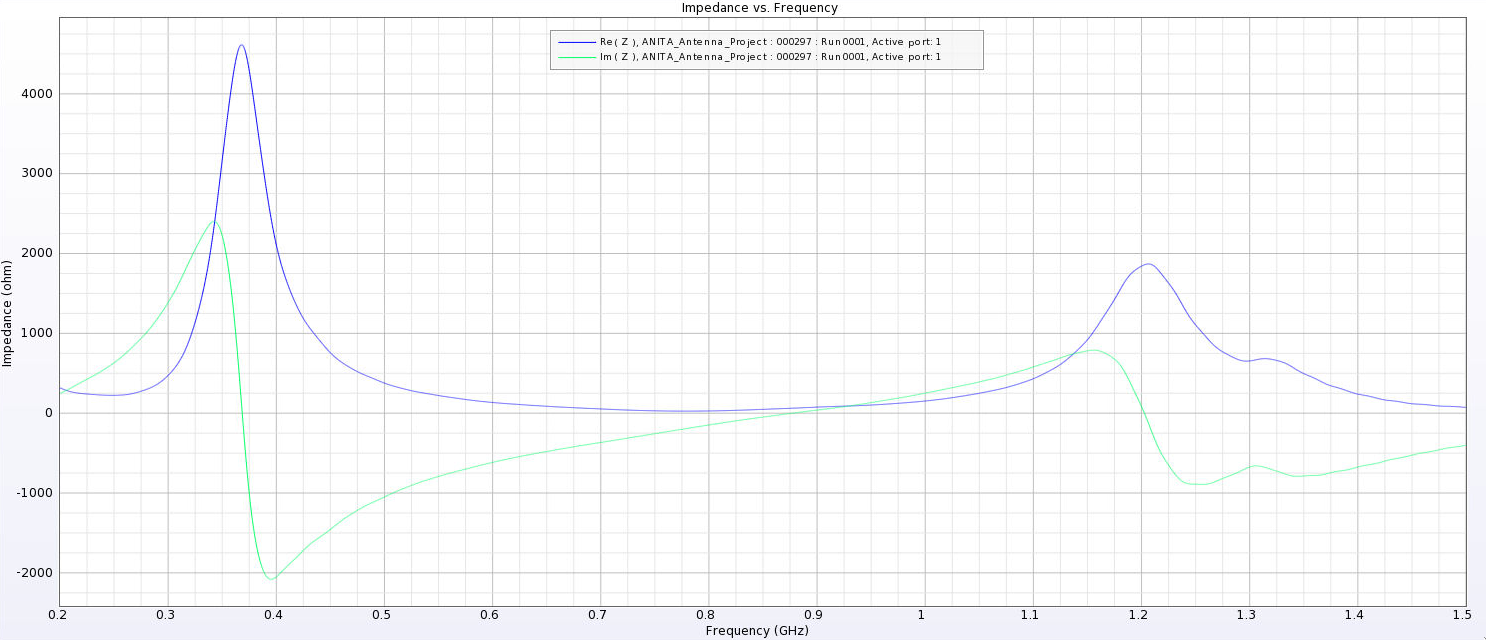 |
| Attachment 9: Separated_Impedance_3_cm.png
| 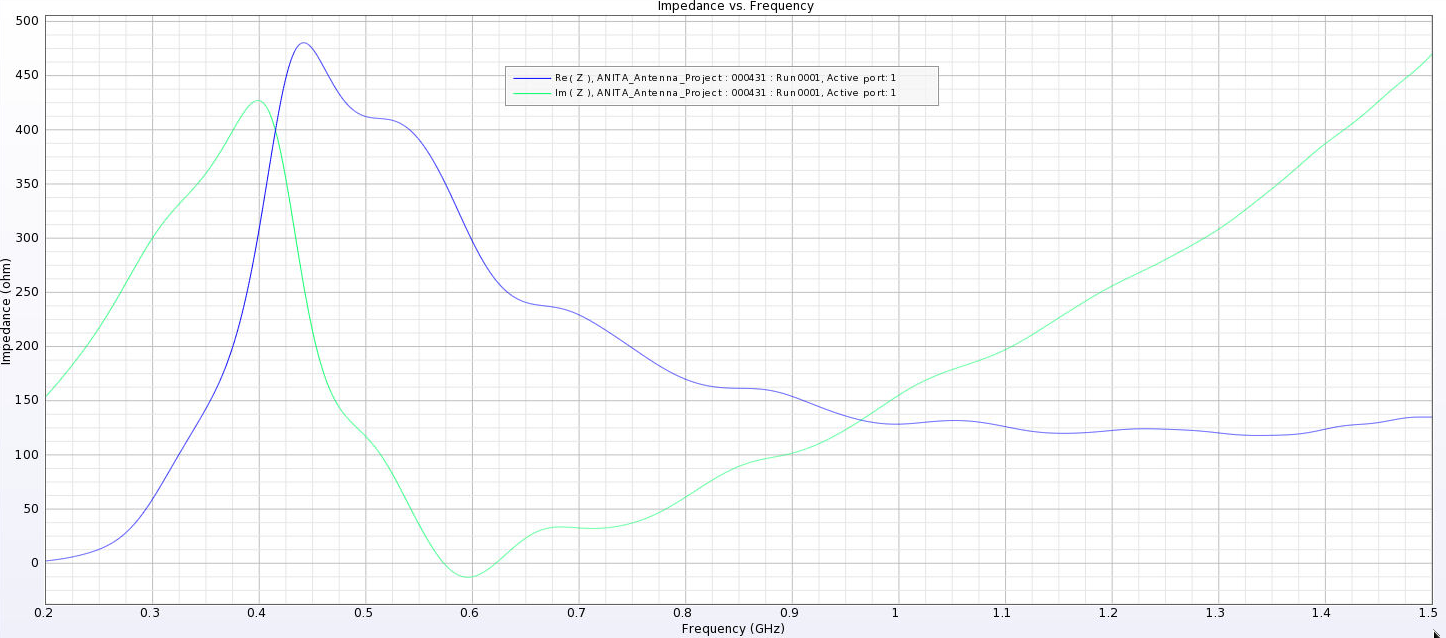 |
| Attachment 10: Separated_Impedance_6_cm.png
| 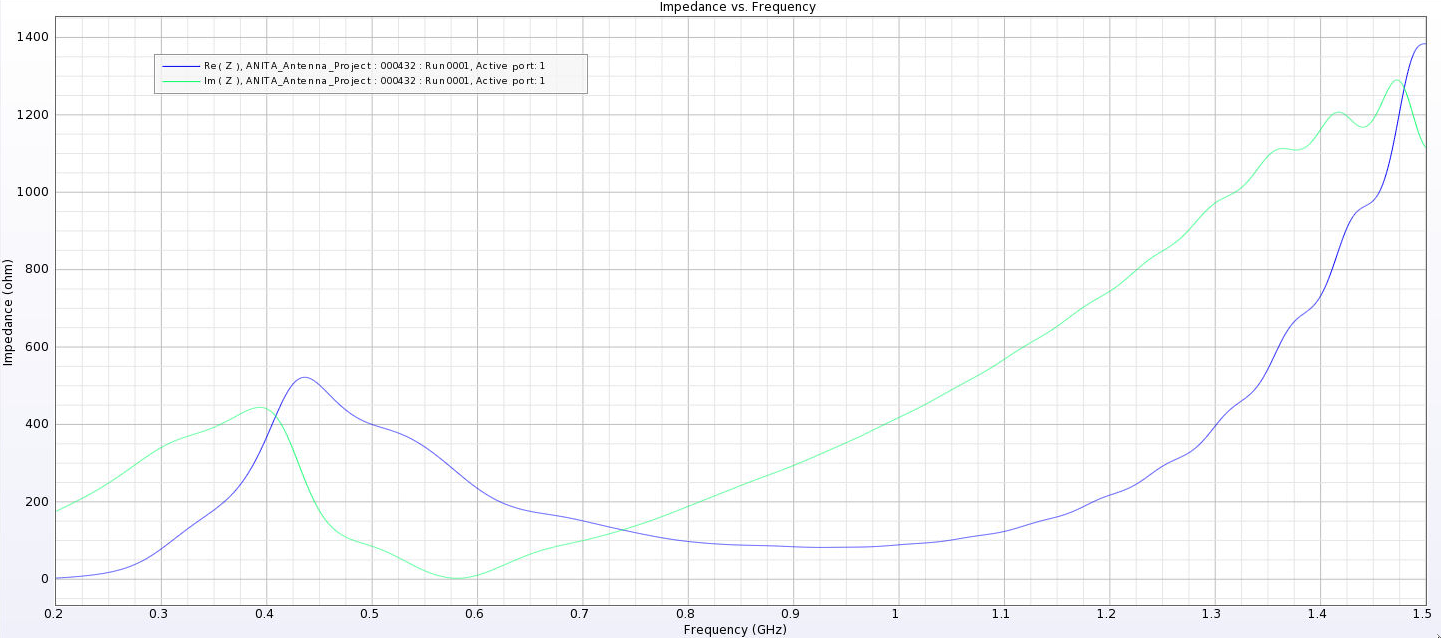 |
| Attachment 11: Separated_Impedance_12_cm.png
| 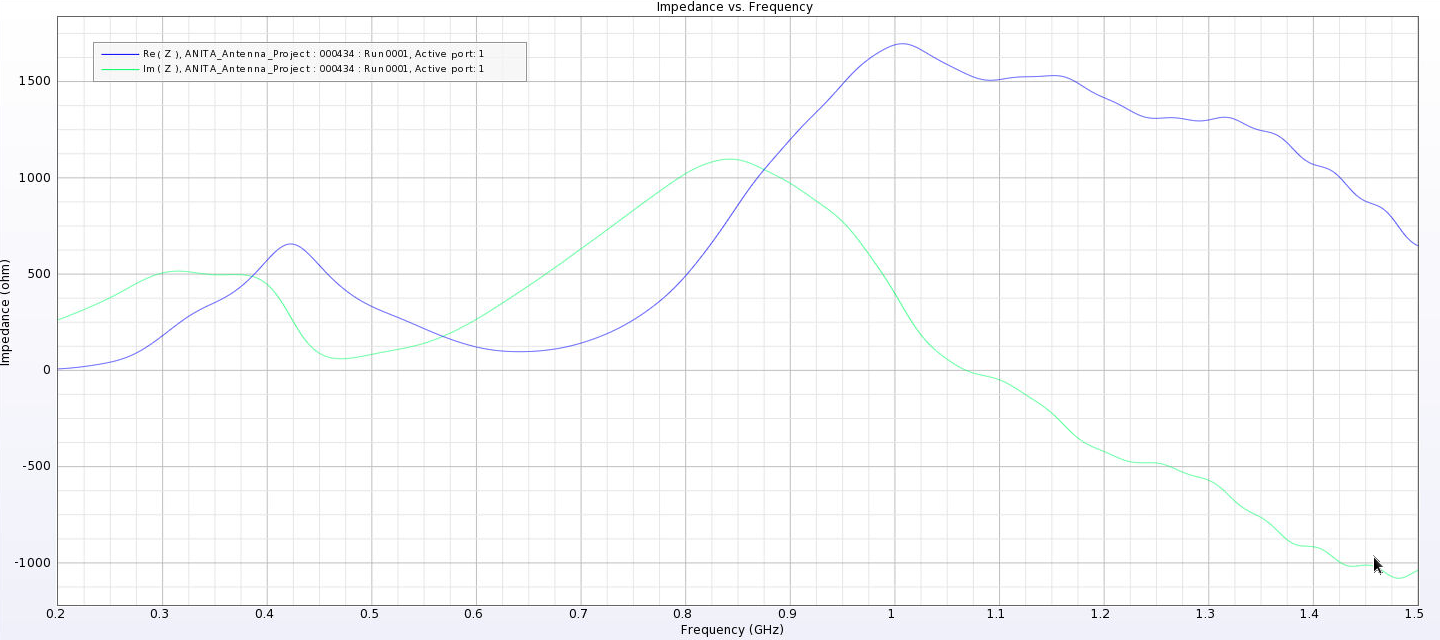 |
| Attachment 12: Separated_Impedance_24_cm.png
| 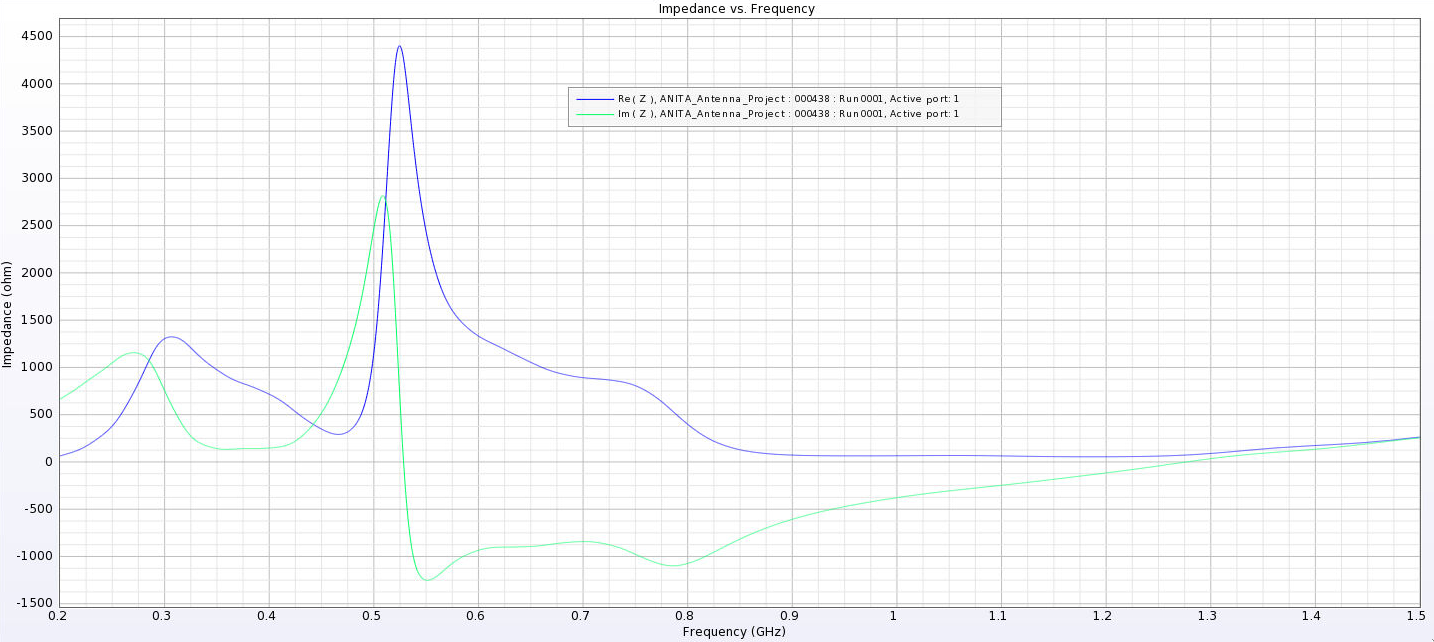 |
| Attachment 13: Separated_Waveguide_Impedance_1.png
|  |
| Attachment 14: Separated_Waveguide_Impedance_3.png
|  |
| Attachment 15: Separated_Waveguide_Impedance_6.png
|  |
| Attachment 16: Separated_Waveguide_Impedance_8.png
|  |
| Attachment 17: Best_Impedance_Plot_Horn_Antenna.png
|  |
|
30
|
Tue Feb 4 17:17:10 2020 |
Julie Rolla | GENETIS working meeting | As of 01/30/2020, the proposal was submitted! The project description and summary are attached below.
Bicone Loop Status Update:
The loop currently runs; however, it does seem that if AraSim jobs are left to run while the user is away, it doesn't properly write one or more of the flags, and makes the loop get stuck at AraSim. We must then re-run from that state by manually editing the Save State text file.
The loop currently has the following additions implemented.
- Functionalized loop (XF_Loop_AraSeed.sh)
- Cleaner bash script which runs smaller bash scripts, instead of one massive script.
- Submits multiple AraSim jobs in parallel for each individual to increase speed.
- Ie if 100k neutrinos are thrown per individual ($NNT=100000, $Seed=10) we can, say, break it up into a "seed" of 10 -- which then submits 10 jobs with NNT=10k for each individual.
- Still needs this to be fixed: The AraSim run in Gen 0 that gets Veff for the ARA actual bicone input file is not currently being seeded. It is just submitting one job the following way.
- If NNT=100000 and Seed=10 (ie 10 jobs with 10k neutrinos), it will only run AraActualBicone as 1 run with 10k neutrino number (ie NNT/seed=NNT_New).
- Save State
- After every "part" -- ie part a, part b, etc -- of the loop, we save state by writing a series of numbers to a text file.
-
If the loop is killed for any reason during the process, the text file is read in, and the loop can continue forward from the last completed item/section.
-
This saves the state after every individual runs through XF.
-
Antenna sizing
-
The larger the antenna, the longer XF takes to run. A true-to-size ARA antenna takes anywhere from 1.5 to 2 hours per individual.
-
What we've done: Allowed us to scale the antenna by a factor of "x".
-
If we make the gaussian centered around something, say, 5 times smaller than the ARA antenna size, we then make the frequencies 5 times larger. This should have the same effect.
-
This also means we've had to scale these sizes back up when plotting.
-
Still needs this to be fixed: all of this has been hardcoded in! We need to make these variables in the MANY places we need to adjust for this scaling. See the to-do list below for details.
Moving forward, I will be doing the following:
- Getting our less coding-savvy teammates familiar with bash, python, and c++. This will be crucial for us when we start adding more parameters to our GA.
- Mastering XF (along with Alex, and Alex).
- Getting the team to clean up some messy coding done during the proposal hustle.
- Fixing some errors.
- Teach the ENTIRE team git.
TO-DO List
The following is our coding to-do list:
-
Get our Ara sim error fixed (Alex, Julie)
-
It seems that if AraSim jobs are left to run while the user is away, it doesn't properly write one or more of the flags, and makes the loop get stuck at AraSim. We must then re-run from that state by manually editing the Save State text file.
-
We are going to try to get XF to run at the command line so that we can submit it as a job. This would alleviate us using an interactive job. We *hope* this would fix the error since the user wouldn't be "gone" while utilizing a GPU in an interactive job.
-
Alex emailed XF to ask a few questions! Stay tuned!
-
Make Hardcoded directories variables.
-
Xmacros (use sed command)
-
All over bash script functions.
-
AraSim job files (sed command?)
-
Make the following variables in the bash script:
In xmacros/simulationPECmacroskeleton_prototype.txt (use sed command):
Grid spacing=0.1
Frequency= factor of n higher
This factor of "n" higher in frequency has to be the same as the Multiplier_factor in the roulette algorithm AND the same as GeoFactor in fitnessFunction_ARA_amy.cpp.
In roulette_algirithm.cpp
Length = 50cm/Multiplier_factor;
STD= 15cm/Multiplier_factor
Radius = 1.5cm/Multiplier_factor;
STD= 0.75cm/Multiplier_factor
Also the "Multiplier_factor" line 89.
This "Multiplier_factor"has to be the same as the GeoFactor in fitnessFunction_ARA_amy.cpp AND the same as the factor higher in frequency in the xmacro.
NOTE:THIS MUST BE RECOMPILED AFTER CHANGING.
In fitnessFunction_ARA_amy.cpp:
GeoFactor=2
This GeoFactor has to be the same as the Multiplier_factor in the roulette algorithm AND the same as the factor higher in frequency in the xmacro
NOTE: THIS MUST BE RECOMPILED AFTER CHANGING.
-
Get rid of all code we are not using anymore.
-
Currently using: XF_loop_AraSeed.sh
-
Look at what items are being called in here, and remove old versions such as XF_loop_Prototype.sh, XF_loop_Functionized.sh, and the related scripts.
-
These can temporarily moved to on "old code" directory and we can delete once we finalize the loop.
-
Turn tournament on in the GA.
-
Parallelize the AraActualBicone job.
-
The AraSim run in Gen 0 that gets Veff for the ARA actual bicone input file is not currently being seeded.
|
| Attachment 1: project_description.pdf
|
| Attachment 2: project_summary.pdf
|
|
4
|
Wed Feb 6 17:36:24 2019 |
Julie Rolla | GENETIS update 2/6 | Atttendance: Julie, Max, Cade, Suren, Evelyn, Sophie
Today's tasks:
Max works on AraSim section (needs to do more research and reading)
Julie, Suren, Sophie, Evelyn work on plotting software to plot gain patterns of indiviudals
Cade preps impuse XF code to be implemented
To-Do list:
I. Divide up sections on paper, and work on them. See http://radiorm.physics.ohio-state.edu/elog/GENETIS/1 for section assignments.
II. Implement impuse code.
- This will require us to change XFintoARA.py so that it takes less output files.
- Is there anything else different with the output files?
III. Finish Evelyn and Sophie's plotting software.
- This means also puting it onto Nutau, and adding in lines to the bash script (both to move and save the images, and to run the code!)
Here are things we can do if we are waiting for Remcom:
I. Change sides of bicone to be sinusoid instead of lines.
II. Take known bicone parameters and put it through AraSim to check if it's working properly.
- This also means we should make plots with it to triple check.
III. Test GA and make it maximize other paramaters to see how it does. |
|